- Coxsackie A viruses (CAVs) are members of the Enterovirus A species in the Picornaviridae family, causing a spectrum from mild mucocutaneous disease to severe neurologic and systemic illness, especially in infants and young children.
- Major contemporary clinical importance relates to CVA6, CVA10 and CVA16 as leading causes of hand-foot-and-mouth disease (HFMD) and herpangina worldwide.
- CVA types can also cause neuroinvasive disease, acute flaccid paralysis, and ocular disease (CVA24 variant keratoconjunctivitis), highlighting diverse tissue tropism.
Taxonomy and Classification
- Family: Picornaviridae
- Genus: Enterovirus
- Species: Enterovirus A (EV-A)
- CAVs within EV-A include multiple serotypes (e.g., CVA2–CVA8, CVA10, CVA12, CVA14, CVA16, CVA24).
Key taxonomic points:
- Grouping as “A” vs “B” was historically based on paralytic phenotype in newborn mice (flaccid vs spastic) but molecular classification now uses VP1 sequence similarity.
- >25 EV-A types are recognized; CVA6 and CVA16 predominate among HFMD-associated types, while CVA10 and CVA24 contribute to HFMD and ocular/neurologic disease.
Structure and Morphology
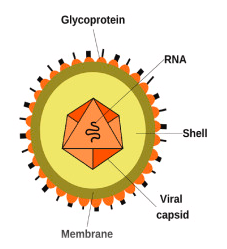
Figure: – Structure and Morphology of Coxsackie A viruses (DOI: https://doi.org/10.1016/j.ab.2023.115368)
- Virion properties
- Small, non-enveloped, icosahedral capsid, ~30 nm in diameter.
- High environmental stability, enabling fecal–oral and respiratory transmission.
- Capsid architecture
Four capsid proteins VP1–VP4:
- VP1, VP2, VP3 on external surface; VP4 internal on the capsid–RNA interface.
- Classical pseudo-T=3 icosahedral symmetry with VP1 around fivefold axes and alternating VP2/VP3 around two- and threefold axes.
Canyon and hydrophobic pocket:
- A surface depression (“canyon”) at the VP1/VP2/VP3 interface often serves as a receptor binding site.
- Underlying VP1 hydrophobic pocket contains a “pocket factor” lipid whose release is linked to uncoating.
- Subtype-specific structural features
CVA10:
- Virion resembles EV-A71 and CVA16 but displays star-shaped VP1 loop protrusions around fivefold axes and distinct VP1 BC/GH loop conformations that shape receptor-binding epitopes.
CVA6:
- Cryo-EM shows a conventional compact virion with VP4 and fatty-acid-like pocket factor; altered (expanded) particles and empty capsids are also produced, but infectious entry is mediated by virions.
Genome Organization and Proteins
Genome:
- Linear, positive-sense ssRNA, ~7.4–7.5 kb, with:
- 5′-UTR containing IRES.
- Single large ORF encoding a polyprotein.
- 3′-UTR and poly(A) tail.
- 5′ end covalently linked to VPg (3B); critical primer for RNA synthesis.
Polyprotein organization (P1–P3):
P1 (structural):
- Cleaved to VP0, VP1, VP3; VP0 further auto-cleaves into VP2 and VP4 in mature virions.
P2 (nonstructural):
- 2A protease (shuts off host cap-dependent translation), 2B and 2C (membrane interactions, replication complex formation).
P3 (nonstructural):
- 3A (membrane anchoring), 3B (VPg), 3C protease, 3Dpol (RNA-dependent RNA polymerase).
Functions:
- Capsid proteins define antigenicity and receptor usage (e.g., VP1 epitopes used for genotyping and neutralization).
- Nonstructural proteins orchestrate replication, host translation shutdown, innate immune antagonism, and cell death.
Replication Cycle
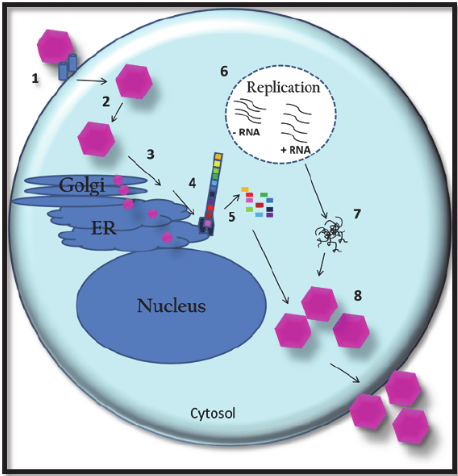
Figure: – Replication Cycle of Coxsackie A viruses (DOI: 10.5772/20932)
Attachment and Entry
Receptor usage (serotype-specific)
- CVA10 and some other EV-A use KREMEN1 as a high-affinity entry receptor, binding via VP1 canyon and loops.
- Other CVA serotypes likely use distinct receptors/co-receptors that are less well defined than for CVB (e.g., CAR, DAF).
Key events during attachment/entry
- High-affinity binding in the VP1 canyon region dislodges a lipid “pocket factor” from the VP1 hydrophobic pocket.
- This triggers conformational changes:
- Externalization of VP4.
- Expansion of the capsid into an “A particle” (uncoating intermediate).
- Virions are internalized by receptor-mediated endocytosis, using clathrin-dependent or other pathways depending on host cell type.
Mechanistic implication: Receptor engagement and pocket-factor ejection mechanically prime the capsid for uncoating, ensuring that genome release is tightly coupled to productive entry.
Uncoating and Genome Release
- Conformational transitions
- Compact virion → expanded A-particle → empty particle plus released RNA.
- Cryo-EM of CVA10 and CVA6 visualizes these discrete structural states, confirming a conserved enteroviral uncoating pathway.
- Genome translocation
- Proton and ion fluxes in endosomes plus receptor-induced structural strain open channels at the two-fold/three-fold axes.
- The positive-sense RNA genome exits through these channels into the cytoplasm, leaving behind an empty capsid.
- Serotype nuance
- For CVA6, intact virions, not pre-expanded particles, are the infectious form entering cells; altered particles and empty capsids are by-products of the cycle, not primary entry vehicles.
Translation and Polyprotein Processing
Immediate translation of genomic RNA
- Released RNA functions directly as mRNA; the 5′ end is linked to VPg (3B) rather than a cap.
- Translation is driven by a type I IRES in the 5′-UTR, recruiting ribosomes in a cap-independent manner.
Polyprotein synthesis and cleavage
- The single ORF is translated into a large polyprotein (P1–P3).
- Viral proteases mediate ordered processing:
2A protease:
- Cleaves the polyprotein at the P1/P2 junction.
- Cleaves host eIF4G, shutting off host cap-dependent translation and shifting ribosomes to viral RNA.
3C protease (and 3CD precursor):
- Further cleaves P2/P3 into mature nonstructural proteins and processes P1 into VP0, VP3, VP1.
Host shutoff and cell remodeling
- Rapid host translation arrest, cytoskeletal changes and stress pathway activation create a translation environment dominated by viral RNA, facilitating high-level protein synthesis and later cytolysis.
Formation of Replication Organelles
- Membrane remodeling
- Nonstructural proteins 2B, 2C, 3A, and their precursors co-opt ER and Golgi membranes, generating vesicular “replication organelles”.
- These structures concentrate:
- Viral RNA and polymerase (3Dpol).
- VPg (3B), 3Cpro, host lipid-modifying enzymes and phosphatidylinositol kinases.
- Autophagy linkage and quasi-envelopment
- CVs (well-studied in CVB, likely similar for CVA) induce and exploit autophagy to:
- Provide double-membrane scaffolds for replication.
- Assist in non-lytic, vesicle-mediated release of quasi-enveloped virions.
- CVs (well-studied in CVB, likely similar for CVA) induce and exploit autophagy to:
- Regulatory RNA structures
- A 5′ cloverleaf RNA interacts with PCBP2 and 3CD to coordinate the switch from translation to replication, acting as a molecular switch shared across enteroviruses, including CV.
RNA Replication
- Initiation
- 3Dpol uses VPg-pUpU as primer to synthesize a full-length negative-strand RNA on the positive template.
- Newly formed negatives serve as templates for multiple positives, amplifying genome copies.
- Kinetics (example CVA16 in RD cells)
- Attachment/uncoating complete by ~1 hour post-infection (p.i.).
- Rapid RNA replication:
- Begins around 3 h p.i., peaking between 3–6 h at MOI 1 and extending to ~9 h at MOI 0.1.
- Viral antigen first detectable at 6 h p.i. (MOI 1) or 9 h p.i. (MOI 0.1), paralleling RNA synthesis kinetics.
- Error-prone polymerase and recombination
- 3Dpol lacks proofreading, generating high mutation rates and allowing frequent recombination; especially in nonstructural regions.
- CVA16, CVA6 and CVA10 show extensive intertypic recombination in P2/P3, influencing fitness and pathogenesis.
Assembly and Maturation
- Capsid assembly
- Structural precursor P1 is processed into VP0, VP1, VP3, which organize into:
- Protomer (VP0–VP1–VP3) → pentamer → empty procapsid.
- Genomic RNA is selectively packaged into procapsids, guided in part by RNA structural motifs that act as packaging signals.
- Structural precursor P1 is processed into VP0, VP1, VP3, which organize into:
- Maturation cleavage
- Within RNA-filled capsids, VP0 auto-cleaves into VP2 and VP4, stabilizing the virion and finalizing maturation.
Release and Spread
- Cytolytic egress
- Classical outcome: rapid cytopathic effect (CPE), culminating in membrane rupture and release of large numbers of virions.
- In CVA16-infected RD cells:
- CPE appears around 12 h p.i., following peak RNA synthesis and virion secretion; secondary infection detectable when virions are released before 9 h p.i. at low MOI.
- Non-lytic vesicular egress
- Like CVB, Coxsackie A viruses can also exit in autophagosome-derived or micro vesicular membranes, yielding quasi-enveloped virions that:
- Evade neutralizing antibodies.
- Spread without overt cell lysis, favoring persistence in some tissues.
- Like CVB, Coxsackie A viruses can also exit in autophagosome-derived or micro vesicular membranes, yielding quasi-enveloped virions that:
- Tissue dissemination
- Local replication at mucosal entry sites (oropharynx, gut) is followed by viremia and seeding of skin, CNS, myocardium, or eye, according to serotype tropism and receptor distribution.
Pathogenesis and Host Immune Response
- Tropism and target organs
- Predominantly infect epithelial cells of oropharynx and intestine, then spread to skin, CNS, muscle, and eye depending on serotype.
- CVA6/CVA10/CVA16: skin and mucosal epithelia (HFMD, herpangina).
- CVA24 variant: ocular surface (acute hemorrhagic conjunctivitis).
- Other CVAs: anterior horn cells, brainstem, leading to acute flaccid paralysis or encephalitis.
- Direct cytopathic effects
- Robust cytolytic replication causes:
- Epidermal cell death and vesicle formation in HFMD.
- Neuronal death in neuroinvasive disease.
- Myocyte damage in some animal models.
- Robust cytolytic replication causes:
- Innate immunity
- Host pattern-recognition receptors (TLR3, MDA5, RIG-I) sense viral RNA, inducing type I IFN and inflammatory cytokines.
- CVA16 proteomic work shows altered expression of numerous host proteins, including HMGB1, and enrichment of cytokine and B-cell receptor pathways, reflecting strong inflammatory activation.
- HMGB1 overexpression reduces CVA16 replication but enhances cytokine release; knockdown increases replication and blunts inflammation, linking HMGB1 to both antiviral defense and immunopathology.
- Adaptive immunity
- Neutralizing antibodies primarily against VP1/VP2/VP3 loops are type-specific; they limit viremia and protect against reinfection but cross-protection between serotypes is limited.
- T-cell responses contribute to clearance but can also amplify tissue injury in CNS and other organs.
Epidemiology and Transmission
- Global distribution
- CVAs are ubiquitous worldwide, with highest burden in low- and middle-income countries and among young children.
- CVA6 and CVA16: leading HFMD pathogens; CVA6 has overtaken EV-A71 and CVA16 in many regions.
- CVA10: increasingly recognized HFMD agent, with multiple genotypes circulating globally and dynamic spatiotemporal patterns.
- CVA24 variant: major cause of acute hemorrhagic conjunctivitis outbreaks.
- Genetic diversity and evolution
- At least 27 CV genotypes; CVA6 and CVA16 show multiple genotypes and recombinant forms, especially in non-structural P2/P3 regions.
- CVA16:
- Five genotypes (G-a to G-e); extensive recombination in non-structural genes while structural proteins remain relatively conserved.
- CVA6:
- Predominant D3 clade with multiple recombinant forms (RF-A, RF-N, RF-H, RF-Y); RF-Y’s 3Dpol region closely related to CVA10, indicating intertypic recombination.
- CVA10:
- Eight VP1-based genotypes (A–H); genotype C dominant in mainland China with distinct regional diffusion routes.
- Transmission routes
- Fecal–oral and respiratory: virus shed in stool, respiratory secretions, and vesicular fluid; incubation ~3–7 days.
- High transmissibility in daycare, schools, and households; seasonal peaks often in warm months.
Clinical Manifestations
- Hand-Foot-and-Mouth Disease (HFMD)
Major etiologic agents
- CVA16 and CVA6 now dominate many HFMD epidemics; CVA10 is an important additional cause; EV-A71 is more often associated with severe neurologic complications.
Typical (“classical”) HFMD
- Cardinal features
- Low- to moderate-grade fever.
- Oral enanthem: painful vesicles/ulcers on tongue, buccal mucosa, soft palate.
- Vesicular/macular rash on palms, soles, occasionally buttocks and genital area.
- CVA16-associated HFMD
- Usually mild, self-limiting; serious neurologic complications occur but are much less frequent than with EV-A71.
CVA6-associated atypical HFMD
- Epidemiologic prominence
- Over the last decade, CVA6 has become a leading HFMD pathogen worldwide, often surpassing CVA16 and EV-A71 in surveillance studies.
- Distinctive clinical features
- Higher fever burden:
- In a Xi’an multicenter cohort (2013–2019; 1,742 CVA6 cases), fever occurred in 82.5% of CVA6 vs 51.3% (EV-A71) and 45.9% (CVA16).
- Rash distribution and morphology:
- More diffuse rash, frequently involving trunk, elbows, knees in addition to classic palmoplantar/oral sites.
- Significantly more polymorphic lesions (macules, papules, vesicles, bullae in the same patient) than in EV-A71/CVA16 HFMD.
- A high rate of bullous rash (20.2% in CVA6 vs <1% in EV-A71/CVA16).
- Sequelae:
- Onychomadesis (nail shedding) occurred in 9.2% of CVA6-infected children in Xi’an, but not in EV-A71 or CVA16 groups.
- Delayed palmoplantar desquamation and nail changes are repeatedly reported in CVA6 outbreaks.
- Higher fever burden:
- Severity spectrum
- Most CVA6 HFMD is mild; however:
- Severe HFMD cohorts show systemic features (prolonged fever, vomiting, diarrhea, elevated CK and blood glucose, positive fecal occult blood) and neurologic complications (e.g., aseptic meningitis) in a minority of cases.
- In one series of 55 severe CVA6 HFMD cases, aseptic meningitis occurred in 52.7% and pulmonary edema in 10.9%.
- Most CVA6 HFMD is mild; however:
CVA10-associated HFMD/herpangina
- Outbreak role
- In a French prospective city-wide study (2010), CVA10 accounted for 39.9% and CVA6 for 28% of HFMD/herpangina cases, together causing two-thirds of infections.
- Clinical pattern
- Frequent herpangina presentation (posterior oral ulcers) often combined with HFMD lesions.
- Fever in ~80% and generally mild disease without neurologic complications in that outbreak.
- Experimental evidence of virulence
- In a neonatal mouse oral infection model, CVA10 caused hind-limb paralysis and death, with high viral loads in muscle, brain, lung and intestine, and neutrophil-dominant inflammation, indicating strong neuro- and myotropism in susceptible hosts.
Herpangina
- Definition and features
- Acute febrile illness with painful vesicles/ulcers on the soft palate, tonsillar pillars, and posterior pharynx, usually without extensive skin lesions.
- Causative CVA serotypes
- Multiple CVA types (CVA2–CVA6, CVA10, CVA16) participate; CVA10 and CVA6 are prominent in modern series.
- Clinical course
- High fever, odynophagia, drooling in toddlers, self-resolving in 3–7 days with supportive care.
Neurologic Disease
Although EV-A71 is the main driver of severe neurologic complications in HFMD, Coxsackie A viruses can also involve the CNS.
- Spectrum of neurologic involvement
- Aseptic meningitis, encephalitis, encephalomyelitis and acute flaccid paralysis (AFP) have been associated with certain CVA serotypes.
- CVA6 severe HFMD
- In the severe CVA6 cohort (55 children), >50% had aseptic meningitis and some developed pulmonary edema, indicating a neurotropic and systemic potential overlapping with, but generally milder than, EV-A71.
- CVA24 and related types
- Prototype CVA24 has been associated historically with poliomyelitis-like AFP; the CVA24 variant (CVA24v), while primarily ocular, underscores how small VP1 changes can re-target CVA to neural vs epithelial tissues.
Ocular Disease (CVA24 Variant)
- Acute hemorrhagic conjunctivitis (AHC)
- CVA24v is a leading cause of AHC, characterized by:
- Sudden onset of painful, intensely red eyes.
- Subconjunctival hemorrhages, chemosis, and watery discharge.
- High contagiosity leading to explosive community outbreaks.
- CVA24v is a leading cause of AHC, characterized by:
- Systemic features
- Usually mild systemic symptoms (fever, malaise); keratitis and visual sequelae are infrequent.
Other / Systemic Manifestations
- Respiratory and gastrointestinal prodromes
- Many CVA infections begin with nonspecific URTI symptoms, low-grade fever, sore throat, and sometimes diarrhea before rash onset.
- Sepsis-like illness in neonates
- Neonatal CVA infections can present with sepsis-like syndromes, myocarditis, hepatitis, or multi-organ failure, though CVB more commonly cause severe neonatal systemic disease.
- Atypical complications
- Case reports describe:
- Otitis media, mastoiditis, and facial nerve palsy after HFMD, illustrating potential for secondary bacterial or inflammatory complications following mucosal damage.
- Case reports describe:
Laboratory and Inflammatory Correlates
- Inflammatory markers in HFMD
- CVA6 HFMD often shows:
- Higher WBC counts than CVA16.
- CRP elevations intermediate between EV-A71 (highest) and CVA16 (lowest).
- CVA6 HFMD often shows:
- Host response in CVA16 infection (cell models)
- Proteomic analysis of CVA16-infected bronchial epithelial cells identified 172 significantly altered proteins, enriched in metabolic, cytokine-receptor, B-cell receptor and neuroactive ligand pathways.
- Up-regulation of HMGB1:
- Overexpression decreased viral replication but increased inflammatory cytokine release.
- Knockdown enhanced replication and reduced cytokines, linking HMGB1 to both antiviral defense and immunopathology.
Laboratory Diagnosis
- Molecular detection
- RT-PCR targeting conserved 5′-UTR or VP1 for pan-enterovirus detection and typing from:
- Throat swabs, stool, vesicle fluid, CSF, conjunctival swabs.
- Sequencing of VP1:
- Gold standard for enterovirus typing and phylogenetic studies (e.g., CVA10, CVA16, CVA6 genotyping).
- RT-PCR targeting conserved 5′-UTR or VP1 for pan-enterovirus detection and typing from:
- Virus isolation
- Cell culture (e.g., RD, HEp-2 cells); some CVA types (e.g., CVA12) show cell-line specific replication kinetics.
- Used in surveillance and research more than routine clinical care.
- Serology
- Neutralization assays and ELISA possible but:
- Type-specific, labor-intensive, and less sensitive in acute settings.
- Mainly for epidemiologic studies or vaccine trials.
- Neutralization assays and ELISA possible but:
Treatment and Antiviral Therapy
- Supportive care
- Mainstay for HFMD and herpangina: hydration, antipyretics, pain control.
- Severe cases (neurologic disease, cardiopulmonary compromise) require intensive support.
- Experimental / broad-spectrum antivirals
- Multiple small-molecule inhibitors of enteroviruses (including CVs) target:
- Capsid pocket (e.g., capsid-binding inhibitors).
- 3Dpol and other nonstructural proteins.
- For CVB, agents like pleconaril and novel inhibitors have shown in vitro/animal efficacy; conceptually relevant to CVAs, but no licensed antivirals specifically approved for CVA.
- Multiple small-molecule inhibitors of enteroviruses (including CVs) target:
- Host-targeted approaches
- Modulation of autophagy, lipid metabolism, and innate immune pathways has been explored in CV infections.
- Proteomic data (e.g., HMGB1 in CVA16) highlight host factors that could serve as antiviral targets.
- Oncolytic virotherapy
- Certain CVs (notably CVA21, CVB3) are being developed as oncolytic agents, exploiting their preference for proliferating cells and CAR/KREMEN1 expression on tumors.
- This is therapeutic use of viruses rather than treatment of natural CVA infection.
Prevention and Control
- Public health and hygiene
- Key measures:
- Hand hygiene, environmental cleaning, cohorting during outbreaks.
- Exclusion of symptomatic children from daycare/school during HFMD or conjunctivitis outbreaks.
- Key measures:
- Vaccines
- Licensed vaccines exist for EV-A71 (and in some regions EV-A71/CVA16 combinations), but no approved monovalent CVA vaccine yet.
- Growing evidence of CVA6/CVA10/CVA16 disease burden has prompted interest in multivalent HFMD vaccines including CVA components.
- Surveillance
- Molecular surveillance of HFMD and enteroviral disease:
- Monitors shift from EV-A71/CVA16 to CVA6/CVA10.
- Tracks emergence of new genotypes/recombinant forms (e.g., CVA6 D3/RF-Y, CVA10 genotype shifts).
- Essential to inform vaccine composition and outbreak control.
- Molecular surveillance of HFMD and enteroviral disease:
Conclusion
Coxsackie A viruses form a diverse group of Enterovirus A types responsible for a major share of HFMD, herpangina, ocular outbreaks, and occasional neuroinvasive disease worldwide. Structurally, they are small, non-enveloped icosahedral virions with a conserved enteroviral genome layout but substantial diversity in VP1-defined antigenic sites and receptor usage. Their pathogenesis reflects a balance between direct cytolytic replication and robust, sometimes damaging host immune responses. While supportive care remains the clinical mainstay and no specific antivirals or CVA-focused vaccines are licensed, rapid advances in molecular epidemiology, structural virology, and host–virus interaction studies are laying groundwork for targeted therapies and multivalent vaccines that include Coxsackie A components.
References
- Machado, R. S., Tavares, F. N., & Sousa, I. P. (2024). Global landscape of coxsackieviruses in human health. Virus Research, 344, 199367. https://doi.org/10.1016/j.virusres.2024.199367
- Zhu, L., Sun, Y., Fan, J., Zhu, B., Cao, L., Gao, Q., Zhang, Y., Liu, H., Rao, Z., & Wang, X. (2018). Structures of Coxsackievirus A10 unveil the molecular mechanisms of receptor binding and viral uncoating. Nature Communications, 9(1). https://doi.org/10.1038/s41467-018-07531-0
- Büttner, C. R., Spurný, R., Füzik, T., & Plevka, P. (2022). Cryo-electron microscopy and image classification reveal the existence and structure of the coxsackievirus A6 virion. Communications Biology, 5(1), 898. https://doi.org/10.1038/s42003-022-03863-2
- Chu, X.-N., Shah, P. T., Ma, Z.-H., Wang, Y., & Xing, L. (2024). Genotyping and phylogeographic dynamics of Coxsackievirus A16. Heliyon, 10(9), e27973. https://doi.org/10.1016/j.heliyon.2024.e27973
- Han, Z., Jia, L., Zhu, R., Fu, H., Lin, C., Huang, H., Deng, L., Zhang, J., & Zhao, L. (2025). The genetic and proliferation characterization analysis of novel coxsackievirus A12 in Beijing, China. Frontiers in Microbiology, 16, 1665461. https://doi.org/10.3389/fmicb.2025.1665461
- Wang, H., Wang, W., Chen, G., Hu, C., Chen, S., Mao, L., & Shen, H. (2024). Molecular characteristics of the structure protein VP1 in Coxsackievirus A10 isolates from China. Journal of Infection in Developing Countries, 18(8), 1025–1034. https://doi.org/10.3855/jidc.18473
- Hu, Y., Liu, C., Yang, J., Zhong, M., Qian, B., Chen, J., Zhang, Y., & Song, J. (2023). HMGB1 is involved in viral replication and the inflammatory response in coxsackievirus A16-infected 16HBE cells via proteomic analysis and identification. Virology Journal, 20(1), 178. https://doi.org/10.1186/s12985-023-02150-8
- Puenpa, J., Saengdao, N., Khanarat, N., Korkong, S., Chansaenroj, J., Yorsaeng, R., Wanlapakorn, N., & Poovorawan, Y. (2022). Evolutionary and genetic recombination analyses of Coxsackievirus A6 variants associated with hand, foot, and mouth disease outbreaks in Thailand between 2019 and 2022. Viruses, 15(1), 34. https://doi.org/10.3390/v15010034
- Cheng, C., Chu, P., Chu, P., Huang, H., Ke, G., Ke, L., Chu, P., & Chu, P. (2024). Phylodynamic and epistatic analysis of Coxsackievirus A24 and its variant. Viruses, 16(8), 1267. https://doi.org/10.3390/v16081267
- Tang, J. W., & Holmes, C. W. (2017). Acute and chronic disease caused by enteroviruses. Virulence, 8(7), 1062–1065. https://doi.org/10.1080/21505594.2017.1308620
- Mustafa, F. H., Ismail, I., Munawar, A. a. Z. A., Basir, B. A., Shueb, R. H., Irekeola, A. A., Ismail, W. Z. W., Jamaludin, J., Balakrishnan, S. R., Sahrim, M., & Yusof, N. Y. (2023). A review on current diagnostic tools and potential optical absorption spectroscopy for HFMD detection. Analytical Biochemistry, 683, 115368. https://doi.org/10.1016/j.ab.2023.115368
- J, P., Walic, M., & S, M. (2011). The key players of Coxsackievirus-Induced myocarditis. In InTech eBooks. https://doi.org/10.5772/20932
