The CRISPR-Cas system consists of a special sequence in prokaryotes, known as clustered regularly interspaced short palindromic repeats (CRISPR) and Cas (CRISPR-associated) genes.

CRISPR was first identified in Escherichia coli in 1987 by Japanese researchers from Osaka University; however, its origin and functions were not initially known. Later, it was found that the original biological function of the CRISPR-Cas system is an RNA-mediated defense system in prokaryotes (bacteria and archaea) against viruses (bacteriophages) or plasmids.
Since their discovery, CRISPR-Cas systems have been transformed into genome editing technology, expanding their applications from research to diagnostics and therapeutics. Essential discoveries in the field of gene editing relating to the CRISPR-Cas9 system also led to the Nobel Prize in Chemistry (2020) for Emmanuelle Charpentier and Jennifer Doudna.
Bacterial Adaptive Immunity of the CRISPR
Bacteria possess several innate immunity-like defenses, such as abortive infection, receptor mutation, and restriction-modification. The CRISPR-Cas system provides a defense mechanism against Mobile genetic elements (MGEs) such as plasmids, bacteriophages, transposons, and insertion sequences.
Although CRISPR sequences were first discovered in bacteria in 1987, their function remained unidentified until 2005, when researchers found that spacer sequences resembled those of bacteriophages. Later, in 2007, it was found that a Streptococcus thermophilus strain acquired resistance against bacteriophages by adding new spacers followed by phage exposure. This finding provided experimental evidence that the CRISPR-Cas system acts as an adaptive immune mechanism in bacteria to defend them against infecting bacteriophages or plasmids.
The CRISPR-Cas system, therefore, works by effectively remembering previous infections by capturing their DNA fragments and later using those memories in the genome to identify and eliminate pathogens based on their sequences.
Structure of the CRISPR Locus
A CRISPR-Cas system consists of a DNA locus called the CRISPR locus, consisting of:
CRISPR array: The CRISPR array consists of short, identical, and palindromic sequences (repeats). Among these repeats, unique spacers derived from foreign DNA fragments (protospacers) are interspersed. These spacers are unique to foreign DNA, are non-repetitive, and act as molecular memories that guide future pathogen recognition. Cas proteins, such as Cas1 and Cas2, primarily mediate spacer integration in the CRISPR array.
Leader Sequence: A variable-length, AT-rich sequence generally adjacent (upstream) to the CRISPR array. The leader sequence typically functions in adaptation of spacers by serving as the landing pad for the Cas1 and Cas2 complex for new spacer integration in the CRISPR locus. These sequences also function as the main promoters for CRISPR pre-crRNA transcription.
CRISPR-associated genes: These genes code for Cas proteins, which are responsible for the integration and elimination of foreign DNA sequences. Cas proteins include Cas1, Cas2, Cas9, Cas12, Cas13 etc. These genes are typically located downstream of the CRISPR-array.
Mechanism of Action of the CRISPR
The CRISPR-Cas system functions in three major steps:
Adaptation (insertion or acquisition):
Adaptation is the process of incorporating a foreign DNA (protospacer) into the CRISPR array as a new spacer sequence, adjacent to the leader sequence. A 2-5-nucleotide-long Protospacer Adjacent Motif (PAM) sequence is located adjacent to the protospacer, which is essential for identifying protospacers. This helps distinguish foreign DNA from the host’s own CRISPR array. The integration of a new spacer is generally mediated in all CRISPR types by the Cas1-Cas2 protein complex.
Expression:
Also called crRNA (CRISPR-RNA) biogenesis, expression is the process of transcription of the new spacer and CRISPR array (containing previous spacers & repeats), and producing a new pre-crRNA. Then, this nascent pre-crRNA is matured with the action of various Cas proteins, such as Cas5 and Cas6, into mature, shorter, and functional crRNA. Each mature crRNA contains a transcribed spacer and transcribed repeat sequences. While the spacer region of the crRNA remains as a linear sequence, the repeats typically form stable hairpins or loop-like structures. These looped structures provide stability by protecting the crRNA from degradation by intracellular nucleases. These structures also provide high-affinity binding for Cas proteins, which bind them to form crRNA-Cas complexes. Later, the crRNA-Cas complex precisely recognizes and binds the complementary protospacer of the foreign DNA.
Interference
During the interference stage, the mature crRNA-Cas protein complex identifies the target foreign DNA molecules. This identification occurs through complementary base pairing of the spacer and protospacer sequence in the presence of Protospacer Adjacent Motifs (PAM) sequences. Here, the crRNA functions to guide the crRNA-Cas complex to the target sequence. After successfully recognizing and binding the target sequence, Cas nucleases, such as Cas9, enable nucleolytic activity, initiating a double-stranded breakage and hence cleaving the target DNA.
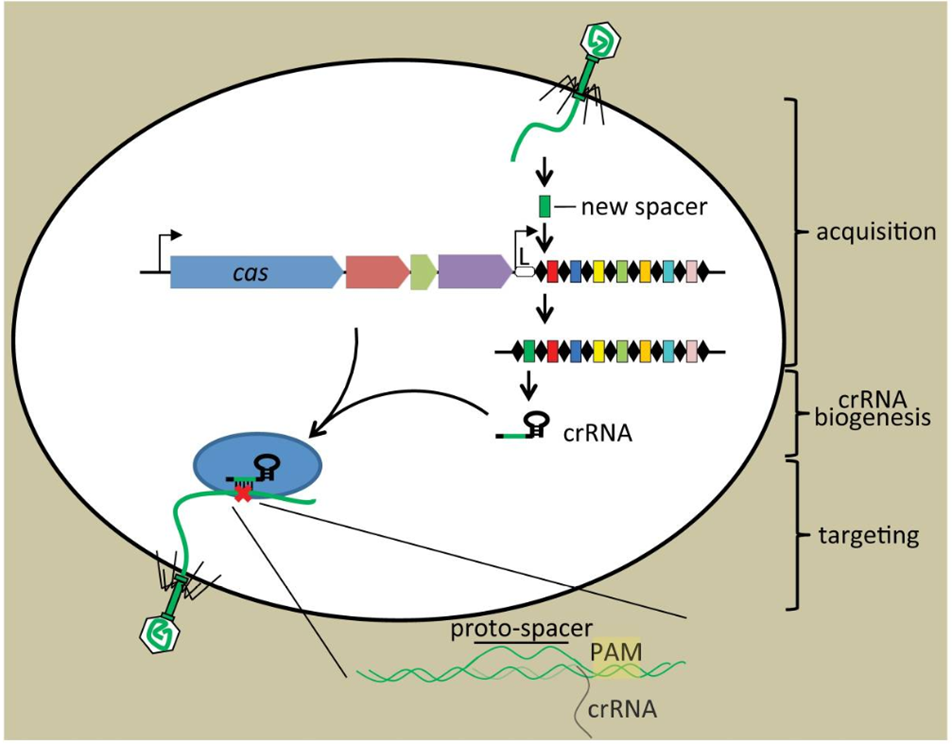
Source: https://doi.org/10.1016/j.molcel.2014.03.011
While the mechanism of action of the CRISPR-Cas system remains universal, specific proteins such as nucleases or crRNA differ according to the variation in different CRISPR-Systems, causing differences in specificity in the action mechanism across all the CRISPR-Cas systems.
Classification of Systems of the CRISPR
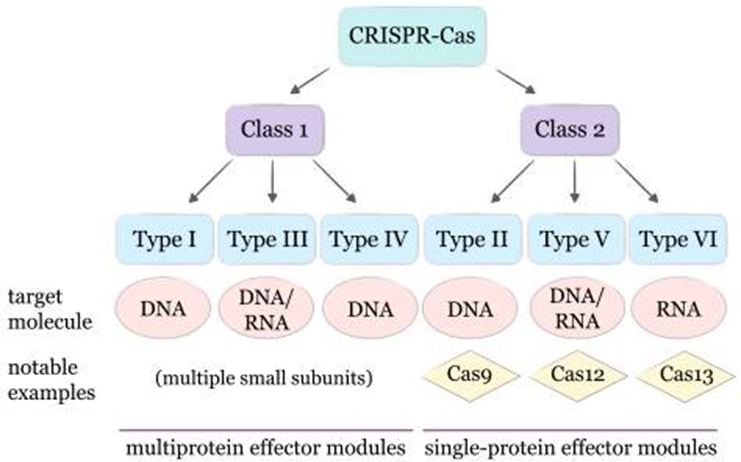
CRISPR-Cas systems are classified into 2 major classes based on the types of their protein complexes. These major classes are further subdivided into 6 primary types and 33 subtypes.
Class1 systems:
Class 1 systems utilize multiprotein effector complexes. These systems are highly prevalent in bacteria and archaea, and primarily target nucleic acids using multiple protein subunits. Type I, Type III, and Type IV are the major types in Class 1 systems.
Type I: Type I CRISPR-Cas systems are the most abundant and widespread CRISPR-Cas systems. The CRISPR-associated complex for antiviral defense (Cascade) complex is mainly responsible for the interference steps in Type I systems. Cascade complex consists of various proteins, such as Cas6, Cas6, Cas7; however, a major protein, Cas3, is responsible for target DNA cleavage.
Type III: Type III CRISPR-Cas systems primarily target RNA. Cas6 protein primarily mediates cleavage of pre-crRNA in this system. A major effector complex in this system is a Csm complex composed of Csm/Cas proteins and crRNA. This system contains 6 subtypes (III-A, III-B, III-C, III-D, III-E, and III-F).
Type IV: Type IV effector complex proteins contain Cas5, Cas7, a unique Cas6, and Csf1 proteins. This system usually lacks Cas proteins during adaptation and cleavage. A unique Cas6 protein is involved in pre-crRNA processing. This system contains 3 subtypes (IV-A, IV-B, and IV-C).
Class 2 systems: Class2 systems utilize a large single-protein effector complex for cleaving target DNA. Type II, Type V, and Type VI are the major types in Class 2 systems.
Type II: In Type II systems, the Cas9 protein with two nuclease domains is associated with the cleavage of the target DNA. Pre-crRNA processing here is mediated by RNase III, trans-activating CRISPR RNA (tracrRNA), and the Cas9 protein. This system contains 3 subtypes (II-A, II-B, and II-C)
Type V: In Type V systems, the Cas12 protein acts as a single effector complex. Cas12 and its domains are responsible for the cleavage of target DNA in this system. This system contains 10 subtypes (V-A, V-B, V-C, V-D, V-E, V-F, V-G, V-H, V-I, and V-K).
Type VI: In Type VI systems, Cas13 protein acts as a single effector complex. The HEPN domain of the Cas13 in type VI systems is involved in cleaving foreign RNA. The type VI system involves 4 subtypes (VI-A, VI-B, VI-C, and VI-D).
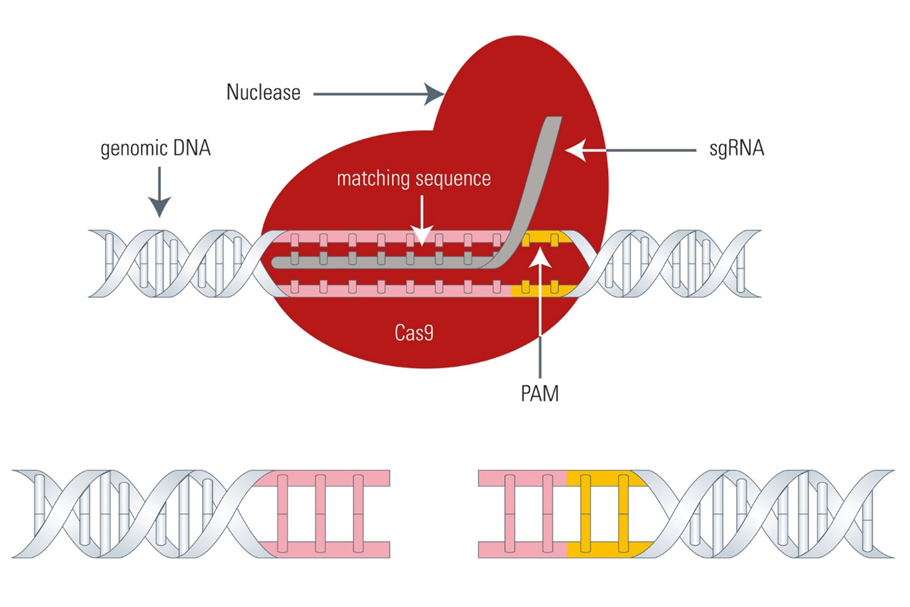
The Cas9 Enzyme of the CRISPR
The Cas9 enzyme is a major component of the bacterial CRISPR-Cas Type II system. It is an RNA-guided enzyme that works to cleave foreign nucleic acid sequences complementary to the RNA loaded into the enzyme during bacterial immunity. It works as molecular scissors to cut DNA at specific locations and hence allows the modification of genomic sequences.
Structurally, the Cas9 protein contains a PAM-interacting domain and two nuclease domains, the NHN and RuvC domains, which work in conjunction with a single guide RNA (sgRNA) and PAM sequence. An active and effective Cas9 complex is composed of Cas9 nuclease, crRNA (CRISPR-RNA) sequence, which specifically targets a particular DNA locus, and tracrRNA, which induces a structural change allowing crRNA to bind to the Cas9 complex, and activates it.
Guide RNA (gRNA) Design of the CRISPR
A guide RNA (also called a single guide RNA; sgRNA) is a synthetic and engineered RNA molecule formed by combining the natural crRNA and tracrRNA, which works to guide the Cas9 nuclease (in the case of CRISPR-Cas9 systems) to a specific target sequence.
While designing a guide RNA, the major goal is targeting efficiency while minimizing or eradicating off-target cleavage. The following fundamentals need to be undertaken:
- Target Region or DNA of interest
Target regions such as introns, exons, or promoters should be defined beforehand. The target region is also dependent upon the approach of the CRISPR-Cas system.
- The Type of Cas protein to be used
Different Cas proteins have different PAM requirements and cleavage modes. The choice of the nuclease also affects the length of gRNA and the expected DNA breakage.
- PAM availability
A PAM sequence must be present next to the target site for the Cas protein to bind and then cleave the DNA. The position and orientation of the PAM determine the exact location where cleavage will occur.
- Testing gRNA for specificity and efficiency
The gRNA should be checked to ensure that it doesn’t bind similar sequences elsewhere than the target region in the genome.
- gRNA GC content and secondary structure
Proper GC content and stable secondary structures of gRNA can increase the efficiency of gRNA pairing with target DNA and its stability for nuclease activity.
Base Editing and Prime Editing of the CRISPR
Base editing and prime editing are the newest versions of the CRISPR-Cas-based technologies, where changes in nucleic acid sequences can be directly introduced without causing double-stranded DNA breakage. Both of these systems use Cas proteins to make a single-stranded nick on the dsDNA.
Base editing converts one DNA base into another (transition mutations) without making double-stranded breakage. Single-nucleotide bases are converted into all four transition mutations (C→T, T→C, A→G, or G→A). In recent times, prime editing has expanded the base-editing platform to all possible transversion mutations (A↔C, A↔T, G↔C, G↔T), along with minor insertions or deletions in the sequence.
Base editing and prime editing are well adapted for large genetic corrections, where vector-mediated delivery of the target gene is constricted due to limitations in gene carrying capacity of viral vectors.
Off-Target Effects and Specificity of the CRISPR
When undesired modifications in DNA sequences occur at sites rather than the target sequence by the action of a CRISPR-Cas complex, causing cleavage of the unintended sequence, it is considered an off-target effect. Off-target effects through CRISPR-Cas can induce insertion or deletion mutations, which can disrupt gene regulation and expression. These lead to undesirable effects, such as oncogenic transformation.
Optimized guide RNA design, use of high-fidelity Cas9 nuclease, and improved delivery methods are potential strategies to reduce off-target effects and improve specificity in CRISPR-Cas gene editing.
Therapeutic Applications of the CRISPR
- Treatment of genetic blood disorders: CRISPR-Cas-based therapies have demonstrated successful clinical trials on hematological conditions, such as sickle cell anemia and β-thalassemia, receiving the first approval from the US-FDA for CRISPR-based therapy.
- Cancer Immunotherapy: CRISPR-Cas-based gene editing is currently applied in inducing multiple knockouts of inhibitory molecules in T-cells, enabling the optimization of CAR-T cells, which are then able to overcome resistance by cancer cells and help the immune system to recognize and attack the tumor cells.
- Treatment of monogenic diseases: CRISPR-Cas systems are being utilized and explored in pre-clinical studies for correcting single-gene mutations through in-vivo genome editing approaches, for disorders such as Duchenne muscular dystrophy (DMD) and cystic fibrosis.
- Selective elimination of supernumerary chromosome: Allele-specific CRISPR-based application focuses on correcting Down syndrome by selectively removing the extra chromosome 21 in trisomic cells.
Ethical and Regulatory Landscape of the CRISPR
- Safety and off-target effects: Unintended off-target mutations leading to massive genomic rearrangements are a significant safety concern.
- Germline Editing Concerns: Genome editing for heritable germlines (eggs, sperms and embryos) remains ethically and socially controversial in terms of consent, long-term effects, and potential use as eugenics rather than disease treatment.
- Distinguishing enhancement with therapy: Differentiating disease treatment from non-medical modifications and enhancement is a major ethical issue, which requires a broad, international scientific agreement.
- Regulatory Bodies: International regulatory organizations such as the US-FDA, the EMA, and the WHO has set-up strict guidelines for CRISPR-based gene therapy involving pre-clinical evaluation, off-target specificity, and research transparency, while emphasizing somatic applications and imposing prohibitions on reproductive genome editing.
Conclusion
CRISPR-Cas systems have changed genome engineering by applying bacterial immunity to transform it into a therapeutic platform. The clinical potentials and applications of CRISPR-based systems continue to expand. However, limitations in specificity, safety, and ethical issues remain important. Responsible therapeutic usage with strong regulation will allow for its long-term impacts on genome editing and next-generation medicine.
References
- Martin Jinek et al., A Programmable Dual-RNA–Guided DNA Endonuclease in Adaptive Bacterial Immunity. Science337, 81621(2012). DOI:10.1126/science.1225829
- Gostimskaya I. (2022). CRISPR-Cas9: A History of Its Discovery and Ethical Considerations of Its Use in Genome Editing. Biochemistry. Biokhimiia, 87(8), 777–788. https://doi.org/10.1134/S0006297922080090
- Liao, C., Sharma, S., Svensson, S.L. et al. Spacer prioritization in CRISPR–Cas9 immunity is enabled by the leader RNA. Nat Microbiol 7, 530–541 (2022). https://doi.org/10.1038/s41564-022-01074-3
- Nidhi, S., Anand, U., Oleksak, P., Tripathi, P., Lal, J. A., Thomas, G., Kuca, K., & Tripathi, V. (2021). Novel CRISPR-Cas Systems: An Updated Review of the Current Achievements, Applications, and Future Research Perspectives. International journal of molecular sciences, 22(7), 3327. https://doi.org/10.3390/ijms22073327
- Koonin, E.V., Makarova, K.S. CRISPR in mobile genetic elements: counter-defense, inter-element competition and RNA-guided transposition. BMC Biol 22, 295 (2024). https://doi.org/10.1186/s12915-024-02090-x
- Omer S Alkhnbashi, Shiraz A Shah, Roger A Garrett, Sita J Saunders, Fabrizio Costa, Rolf Backofen, Characterizing leader sequences of CRISPR loci, Bioinformatics, Volume 32, Issue 17, September 2016, Pages i576–i585, https://doi.org/10.1093/bioinformatics/btw454
- Barrangou, R., & Marraffini, L. A. (2014). CRISPR-Cas systems: Prokaryotes upgrade to adaptive immunity. Molecular Cell, 54(2), 234–244. https://doi.org/10.1016/j.molcel.2014.03.011
- Joseph, C. E., Jain, A., Yaqub, M. O., & Edison, L. K. (2025). CRISPR-Cas Systems: Bridging Bacterial Immunity and Host Interactions. Applied Microbiology, 5(4), 118. https://doi.org/10.3390/applmicrobiol5040118
- Navarro, C., Díaz, M. P., Durán, P., Castro, A., Díaz, A., Cano, C., Carbonell-Zabaleta, A.-K., Solano-Jiménez, D.-S., Rivera-Porras, D., Contreras-Velásquez, J. C., & Bermúdez, V. (2025). CRISPR-Cas Systems: A Functional Perspective and Innovations. International Journal of Molecular Sciences, 26(8), 3645. https://doi.org/10.3390/ijms26083645
- Xu, Z., Chen, S., Wu, W., Wen, Y., & Cao, H. (2023). Type I CRISPR-Cas-mediated microbial gene editing and regulation. AIMS microbiology, 9(4), 780–800. https://doi.org/10.3934/microbiol.2023040
- Mir, A., Edraki, A., Lee, J., & Sontheimer, E. J. (2018). Type II-C CRISPR-Cas9 Biology, Mechanism, and Application. ACS Chemical Biology, 13(2), 357–365. https://doi.org/10.1021/acschembio.7b00855
- Kantor, A., McClements, M. E., & MacLaren, R. E. (2020). CRISPR-Cas9 DNA Base-Editing and Prime-Editing. International journal of molecular sciences, 21(17), 6240. https://doi.org/10.3390/ijms21176240
- Kalter, N., Fuster-García, C., Silva, A., Ronco-Díaz, V., Roncelli, S., Turchiano, G., Gorodkin, J., Cathomen, T., Benabdellah, K., Lee, C., & Hendel, A. (2025). Off-target effects in CRISPR-Cas genome editing for human therapeutics: Progress and challenges. Molecular Therapy: Nucleic Acids, 36(3), 102636. https://doi.org/10.1016/j.omtn.2025.102636
- https://www.fda.gov/news-events/press-announcements/fda-approves-first-gene-therapies-treat-patients-sickle-cell-disease
- Feng, X., Li, Z., Liu, Y., Chen, D., & Zhou, Z. (2024). CRISPR/Cas9 technology for advancements in cancer immunotherapy: from uncovering regulatory mechanisms to therapeutic applications. Experimental hematology & oncology, 13(1), 102. https://doi.org/10.1186/s40164-024-00570-y
- Ryotaro Hashizume, Sachiko Wakita, Hirofumi Sawada, Shin-ichiro Takebayashi, Yasuji Kitabatake, Yoshitaka Miyagawa, Yoshifumi S Hirokawa, Hiroshi Imai, Hiroki Kurahashi, Trisomic rescue via allele-specific multiple chromosome cleavage using CRISPR-Cas9 in trisomy 21 cells, PNAS Nexus, Volume 4, Issue 2, February 2025, pgaf022, https://doi.org/10.1093/pnasnexus/pgaf022
