Human rhinoviruses (HRVs) are the leading cause of the common cold but also play major roles in severe respiratory disease and asthma exacerbations, especially in children and high-risk adults.

- Human rhinoviruses (HRVs) are:
- The most frequent viral cause of upper respiratory infections worldwide.
- Increasingly recognized as important pathogens in lower respiratory tract disease, wheezing, asthma, bronchiolitis, and COPD exacerbations.
- Historically viewed as mild, HRVs are now linked to:
- Hospitalizations, especially in infants, the elderly, immunocompromised, and those with chronic lung disease.
- Long-term respiratory morbidity, including recurrent wheeze and asthma development after early-life HRV bronchiolitis.
- No licensed antiviral or vaccine currently exists, despite the heavy global burden.
HRVs also serve as important models for enterovirus biology and for understanding viral interference with influenza and SARS-CoV-2.
Taxonomy and Classification
- Family: Picornaviridae
- Genus: Enterovirus
- Species:
- HRV-A, HRV-B, HRV-C.
- Key points:
- Reclassified from a separate “rhinovirus” genus into Enterovirus based on genomic and structural similarity.
- 160–170 genotypes across the three species.
- HRV-A and HRV-C are most often associated with severe lower airway illness and wheezing; HRV-B is generally less virulent.
Structure and Morphology
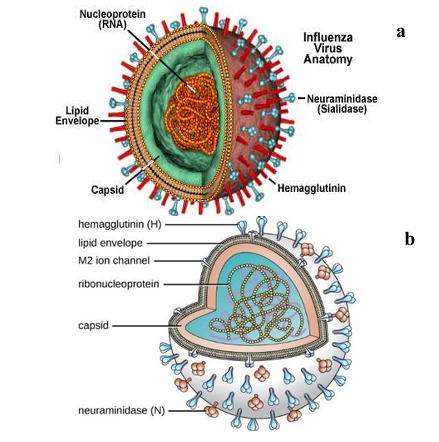
Fig 1: Anatomy of Human Rhino Virus (HRV) (Khan et al., 2021)
- Virion properties:
- Small (~30 nm), non-enveloped, icosahedral capsid.
- Capsid composed of 60 copies each of VP1, VP2, VP3, VP4.
- VP1, VP2, VP3 constitute the outer surface, determining antigenic diversity; VP4 is internal and anchors RNA to the capsid.
- Canyon structure and receptors:
- VP1 forms a surface “canyon” around the five-fold axis that acts as the receptor-binding site.
- Many HRV-A/B types use ICAM-1; some HRV-A use LDL receptor family; HRV-C uses CDHR3 (especially asthma-associated variants).
- Morphological simplicity vs. complexity of diversity:
- Lack of lipid envelope → environmental stability, resistance to desiccation, and many disinfectants.
- High heterogeneity in capsid proteins creates extensive serotypic diversity, complicating vaccine design.
Genomic Organization and Proteins
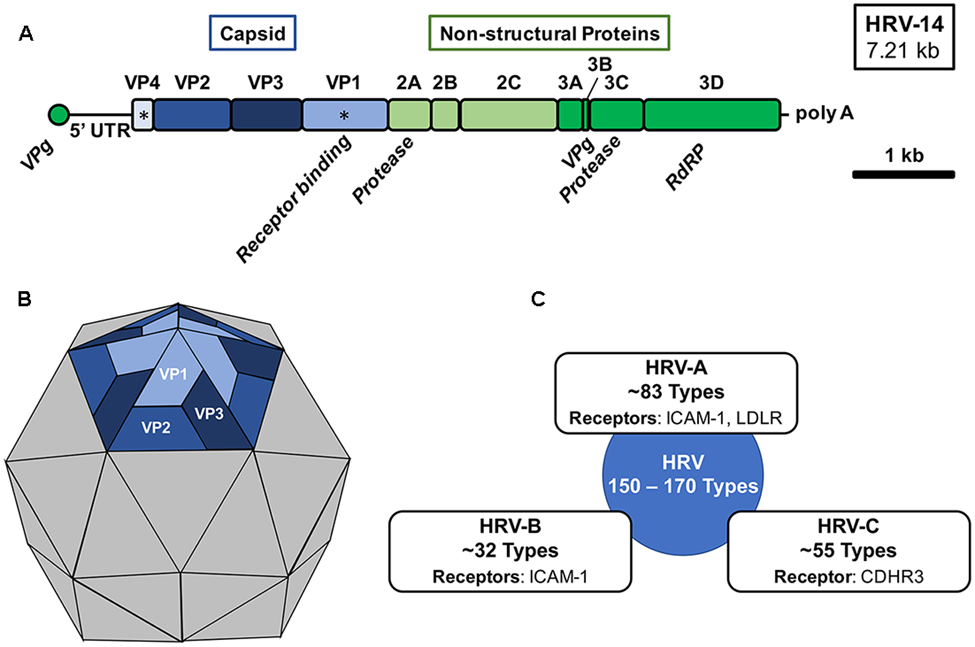
Fig 2: Human rhinovirus genomic organization, virion structure, and species (Stobart et al., 2017)
- Genome overview
- Human rhinoviruses (HRVs) are positive-sense, single-stranded RNA viruses, ~7.2–7.3 kb long.
- Genome is non-segmented and functions directly as mRNA once uncoated.
- 5′ end: covalently linked VPg protein, plus a long structured 5′ untranslated region (5′UTR) containing an internal ribosome entry site (IRES).
- 3′ end: short 3′UTR followed by a poly(A) tail.
- Genome layout
- Single large open reading frame (ORF) encodes one polyprotein, later cleaved into structural and non-structural proteins.
- Genomic organization: 5′UTR–P1–P2–P3–3′UTR–poly(A).
- P1: capsid (structural) proteins VP4, VP2, VP3, VP1.
- P2 and P3: non-structural proteins 2A, 2B, 2C, 3A, 3B(VPg), 3C, 3D.
- 5′UTR and IRES
- 5′UTR is hypervariable in some regions but retains conserved structural elements for type I IRES function.
- IRES mediates cap-independent translation, allowing viral protein synthesis even when host cap-dependent translation is shut down.
- Full-genome sequencing of 99 HRV reference strains showed a common IRES configuration favoring rapid, nonscanning initiation of translation.
- Polyprotein processing
- Polyprotein is co- and post-translationally cleaved by viral proteases 2Apro and 3Cpro/3CD into 11–12 mature proteins.
- Intermediate cleavage products also participate in replication complex formation.
- Structural (capsid) proteins
- HRV capsid: 60 copies each of VP1, VP2, VP3, VP4 arranged in icosahedral symmetry.
- VP1–VP3 form the outer shell and determine antigenicity and receptor binding; VP4 lines the interior and anchors the RNA.
- High-resolution structures (e.g., HRV16, HRV2, HRV14) show:
- A “canyon” around each five-fold axis on VP1/VP2/VP3 that binds receptors like ICAM-1.
- A hydrophobic pocket in VP1 occupied by a “pocket factor”; binding of capsid-targeting drugs here stabilizes the capsid and blocks uncoating.
- Recent cryo-EM work on HRV B14/HRV2 revealed that ~12% of the RNA forms 13-bp dsRNA duplexes organized into a dodecahedral cage attached to basic residues on the inner capsid; this cage coordinates assembly and controls uncoating timing.
- Non-structural proteins and key functions
- 2Apro: cysteine protease that cleaves host eIF4G, inhibiting cap-dependent host translation and privileged IRES-driven viral translation.
- 2B/2BC, 3A: small hydrophobic proteins that remodel intracellular membranes and help form replication organelles.
- 2C: NTPase/helicase-like protein integral to the replication complex.
- 3B (VPg): primer for RNA synthesis and linked to the 5′ end of viral RNA.
- 3Cpro: main viral protease processing the polyprotein; also cleaves host factors, modulating immune responses.
- 3Dpol: RNA-dependent RNA polymerase with low fidelity, generating quasispecies and supporting extensive diversity among HRV types.
- Genomic diversity and classification
- Full-genome analysis identified three major species (HRV-A, HRV-B, HRV-C) and a candidate clade “HRV-D”.
- Diversity spans capsid genes, 5′ spacer elements, protease regions, and 3′UTRs, helping explain clinical heterogeneity and variable responses to antivirals.
Replication Cycle
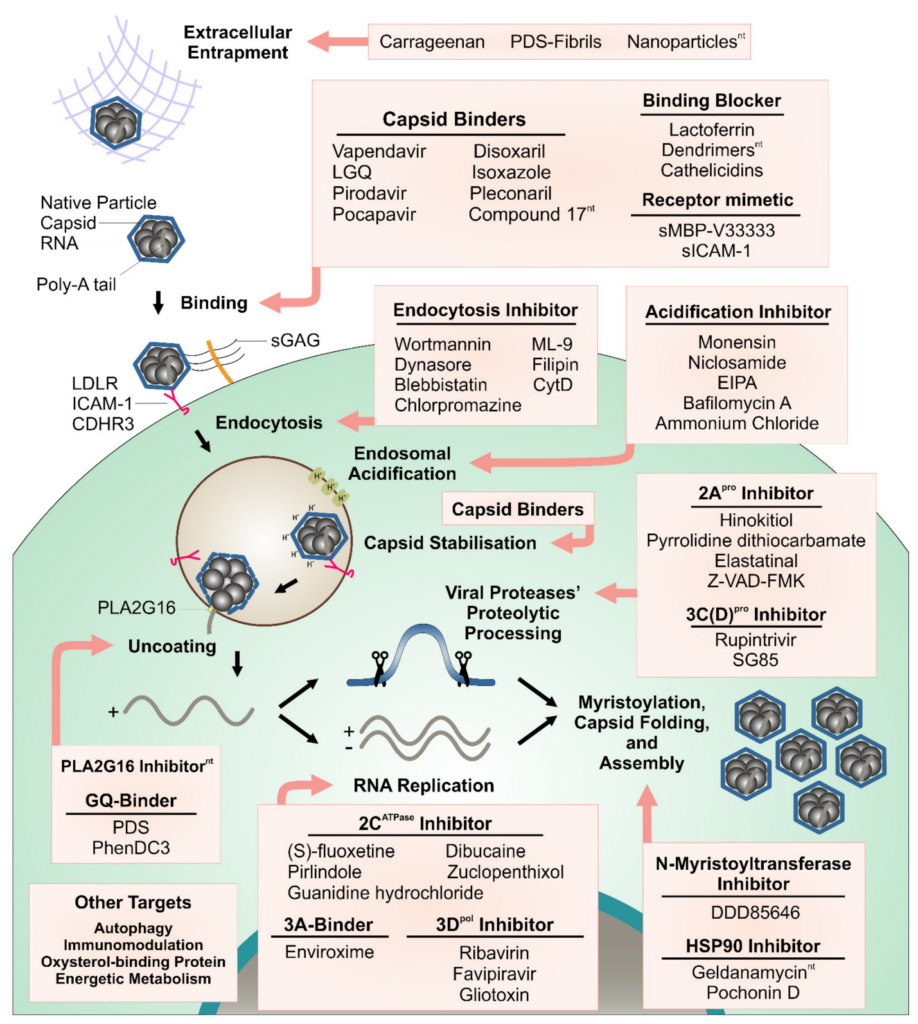
Fig 3: Steps of the rhinovirus replication cycle targeted with antiviral compounds (Real-Hohn & Blaas, 2021)
- Attachment
- HRVs selectively infect ciliated airway epithelial cells in nasal and bronchial mucosa.
- Receptors differ by species/serotype:
- Major group HRV-A/B: ICAM-1 (intercellular adhesion molecule-1).
- Some minor-group HRV-A: LDL receptor family.
- HRV-C: CDHR3, where certain variants (e.g., CDHR3-Y529) increase susceptibility and asthma risk.
- Entry and uncoating
- After receptor binding, virions are internalized via endocytic pathways; mild acidic pH and/or receptor engagement trigger structural rearrangements.
- Structural changes include:
- Hinge movement of VP1 around its hydrophobic pocket.
- Coordinated displacement of VP2 and VP3.
- Capsid expansion and opening of channels at two-fold axes and at canyon bases, serving as gates for VP1 N-terminus externalization and RNA extrusion.
- Capsid “breathing” allows transient exposure of internal peptides (VP4, VP1 N-termini) that can interact with membranes and facilitate genome release.
- RNA exists as a continuous strand through a ~10 Å channel, leaving behind an empty 80S-like capsid. Capsid-RNA duplex cage interactions help prevent premature uncoating while enabling efficient release once triggered.
- Translation
- Positive-sense RNA is directly translated by host ribosomes using the IRES; no 5′ cap is required.
- Early translation produces polyprotein intermediates and proteases 2A and 3C, which shut off host translation (via eIF4G and PABP cleavage) and promote viral takeover.
- Non-structural proteins recruit host membranes and factors to form replication complexes.
- RNA replication
- Replication occurs on virus-induced membrane vesicles derived from the ER/Golgi.
- 3Dpol uses VPg-primed positive-strand RNA to synthesize a negative-strand template, then multiple positive strands, resulting in ~10,000-fold more positive than negative strands in airway epithelial cells.
- In differentiated human bronchial epithelial (HBE) cultures, viral RNA levels plateau at relatively constant values, and infection is restricted to a small fraction of cells, with minimal overt cytopathic effect.
- Assembly and maturation
- Structural proteins assemble into protomers → pentamers → icosahedral capsids that package RNA.
- RNA–capsid interactions, including the dodecahedral dsRNA cage, guide correct assembly and ensure the genome is ordered at the inner capsid surface.
- Maturation includes rearrangements in VP0/VP4/VP2 and stabilization of the hydrophobic pocket.
- Release and spread
- HRVs are mostly released by cell lysis, especially in standard cell lines, but in primary airway epithelia, infection can be non-lytic, with limited cell death and continued epithelial integrity.
- Virions spread locally along the airway epithelium and via secretions into the upper airway, facilitating droplet and contact transmission.
Pathogenesis and Host Immune Response
- Primary infection site and cell targets
- Main targets: nasal and bronchial ciliated epithelial cells; infection is typically focal, affecting only a small percentage of cells in differentiated cultures.
- HRVs are traditionally viewed as upper airway pathogens, but HRV-A and HRV-C also efficiently infect lower airways, particularly in individuals with asthma or COPD.
- Host recognition of HRV
- Viral RNA is sensed as pathogen-associated molecular patterns (PAMPs):
- Endosomal TLR3, TLR7/8 recognize dsRNA and ssRNA.
- Cytosolic RIG-I and MDA5 recognize viral RNA replication intermediates.
- Activation of these pattern-recognition receptors triggers downstream transcription factors (NF-κB, IRF3/7) and induction of type I and type III interferons (IFN-α/β, IFN-λ) and multiple cytokines.
- Viral RNA is sensed as pathogen-associated molecular patterns (PAMPs):
- Innate antiviral responses
- In nasal and bronchial epithelial models, HRV infection strongly induces type I and III IFNs, with expression levels modulated by viral dose.
- IFNs stimulate expression of interferon-stimulated genes (ISGs) such as viperin and ISG15, which can be induced partly independently of IFN levels, suggesting direct PRR–ISG pathways.
- In human lung slices, HRV induces robust IFN-α2a, IFN-γ, CXCL10, CXCL11, CCL5, TNF-α, reflecting strong antiviral and inflammatory responses.
- Epithelial-driven inflammation and asthma/COPD links
- HRV-infected epithelium secretes chemokines IL-8 (CXCL8), IL-6, and others that recruit neutrophils, eosinophils, monocytes, and T cells to the airway.
- In asthma and COPD:
- HRV infection triggers epithelial damage, mucus hypersecretion, airway obstruction, and gene expression changes resembling chronic disease signatures.
- Lung tissue models show RV-induced gene expression overlaps with COPD/asthma signatures in epithelial ciliation, barrier, and inflammatory pathways.
- Immune evasion and dysregulation
- HRV non-structural proteins (2A, 3C, 3A, others) inhibit host protein synthesis and interfere with IFN signaling pathways (as known for picornaviruses).
- In asthma, HRV can both induce and subvert interferon responses:
- Deficient or delayed type I IFN in some asthmatics may permit higher viral replication.
- HRV also upregulates immunosuppressive mediators (e.g., TGF-β, PD-L1, IDO, LAP3) that dampen antiviral immunity and favor persistent inflammation and remodeling.
- Why disease severity varies
- Many HRV infections are asymptomatic or mild; symptom severity does not always correlate with viral load or cytopathic effect.
- A key determinant appears to be host response intensity, particularly the magnitude and pattern of cytokine and interferon responses, rather than direct tissue destruction.
- Children with atopy or asthma and adults with chronic lung disease show exaggerated inflammatory responses and are prone to wheezing and severe exacerbations during HRV infection.
Epidemiology and Transmission
- Burden and prevalence:
- HRVs are detected in 13–59% of respiratory tract infections globally, often the most common viral agent.
- In a Croatian pediatric dataset, HRVs accounted for 33.4% of viral respiratory infections, higher than adenoviruses, seasonal coronaviruses, and bocaviruses.
- Seasonality:
- Often peaks in autumn and spring, but circulation can be year-round and less sharply seasonal than influenza.
- During COVID-19, HRV circulation was less suppressed than influenza and RSV, despite public health measures.
- Transmission routes:
- Mainly via respiratory droplets and aerosols, and fomite contact with contaminated secretions.
- Non-enveloped stability enhances survival on surfaces and in the environment.
- Population at risk:
- Children experience the highest infection rates and are major drivers of community transmission.
- HRVs are key triggers of asthma exacerbations and COPD flare-ups in both children and adults.
Clinical Manifestations
- Incubation period and general features
- Incubation: typically, 1–3 days after exposure.
- Duration: symptoms usually peak around day 2–3 and resolve within 7–10 days, though cough may persist longer.
- Typical “common cold” presentation
- Most frequently observed manifestations in outpatients with acute upper RTIs:
- Nasal discharge, nasal congestion, runny nose.
- Cough and sore throat.
- Mild headache, malaise, and occasionally low-grade fever (fever less common than in influenza).
- In a large comparative cohort (849 HRV vs 417 influenza patients), cough, nasal discharge, and sore throat predominated in HRV, whereas fever and systemic symptoms were more prominent in influenza.
- Most frequently observed manifestations in outpatients with acute upper RTIs:
- Lower respiratory tract involvement
- HRV can cause:
- Bronchiolitis and pneumonia, especially in infants, the elderly, and immunocompromised patients.
- Viral wheeze and asthma or COPD exacerbations; HRV is implicated in up to two-thirds of exacerbations in children and over half in adults.
- HRV infection in human lung tissue induces gene signatures mirroring COPD and asthma, supporting a mechanistic role in disease worsening.
- HRV can cause:
- Disease severity and risk factors
- Most infections are mild, but severe acute respiratory symptoms occur more frequently in:
- Patients with asthma, allergic rhinitis, and diabetes mellitus, especially when infected with HRV-C.
- Elderly individuals and those with cardiac or chronic pulmonary conditions.
- In HRV-C-infected outpatients, coexisting diabetes (aOR 3.6) and asthma (aOR 1.9) significantly increased the odds of severe acute respiratory symptoms; asthma and allergic rhinitis strongly predicted recurrent RTI-related primary care visits over 1 year.
- Most infections are mild, but severe acute respiratory symptoms occur more frequently in:
- Complications
- Upper airway: otitis media, sinusitis, acute rhinosinusitis, especially in children.
- Lower airway:
- Severe bronchiolitis and hospitalization in infants.
- Pneumonia and respiratory failure in immunocompromised or frail adults.
- Chronic consequences: early-life HRV-associated wheeze illnesses are linked to increased risk of recurrent wheeze and asthma development later in childhood.
- Clinical differentiation from other respiratory viruses
- Compared with influenza:
- HRV infections show less frequent high fever, myalgia, and severe systemic symptoms but similar overall burden in high-risk patients.
- During the COVID-19 era, HRV circulation continued robustly despite public health measures, highlighting its efficient transmission and role as a persistent cause of cold-like and asthma-exacerbating illness.
- Compared with influenza:
Laboratory Diagnosis
- Specimens:
- Nasopharyngeal swabs, aspirates, or washes are preferred.
- Lower respiratory specimens (BAL, sputum) in severe or ICU cases.
- Molecular methods (standard of care):
- RT-PCR or real-time RT-PCR targeting conserved 5′UTR for detection.
- Multiplex respiratory panels commonly include HRV/enterovirus as a combined or separate target, allowing simultaneous detection of multiple viruses.
- Genotyping based on VP4/VP2 or VP1 sequences for epidemiology and species assignment.
- Other methods:
- Classical viral culture is now rarely used (slow, less sensitive).
- Antigen tests are generally less sensitive than nucleic acid amplification for HRV.
- Diagnostic challenges:
- High rate of asymptomatic carriage and prolonged post-symptomatic shedding complicate interpretation, particularly in high-risk pediatric populations.
- Co-detections with other viruses are common; clinical significance must be interpreted in context.
Treatment and Antiviral Therapy
- Current clinical management:
- Largely supportive: hydration, antipyretics, symptomatic relief.
- Management of underlying conditions (e.g., bronchodilators and steroids for asthma/COPD exacerbations).
- No approved HRV-specific antiviral drug.
- Investigational and candidate antivirals:
- Capsid-binding agents (e.g., pleconaril-like drugs):
- Bind a hydrophobic pocket in VP1, stabilize the capsid, and interfere with uncoating.
- Showed antiviral activity, but many candidates failed due to limited efficacy or side effects.
- Protease inhibitors:
- Target 3C^pro or 2A^pro, blocking polyprotein processing; conceptually promising given the success of protease inhibitors against other viruses.
- Polymerase and replication complex inhibitors:
- Small molecules targeting 3D^pol or host factors involved in replication organelle formation.
- Host-targeted therapies:
- Inhibitors of host pathways needed for attachment or replication (e.g., receptor blockade, lipid metabolism).
- RNA-targeting strategies:
- Stabilization of G-quadruplex or other RNA structures to interfere with genome uncoating and replication has shown in vitro promise.
- Capsid-binding agents (e.g., pleconaril-like drugs):
- Azithromycin and other repurposed drugs:
- Azithromycin has demonstrated antiviral and immunomodulatory effects against enteroviruses and HRVs in experimental systems, including modulation of interferon responses, but its clinical benefit for HRV is unproven, and use is limited by resistance concerns.
Prevention and Control
- General public health and behavioral measures:
- Hand hygiene and surface disinfection (though HRV is relatively resistant to some disinfectants).
- Respiratory etiquette and avoidance of close contact when symptomatic.
- These interventions variably reduce HRV spread; during COVID-19, they were less effective against HRV than against influenza/RSV, likely due to HRV’s stability and efficient transmission.
- Vaccination – current status and challenges:
- No licensed HRV vaccine; development has been hindered by:
- Large number of serotypes and limited cross-reactive neutralizing antibodies.
- Incomplete understanding of correlates of protection, especially at mucosal surfaces.
- Recent progress:
- Structural and immune profiling have identified conserved B-cell epitopes on capsid proteins and conserved T-cell epitopes, encouraging high-valent or epitope-focused vaccines.
- Multi-epitope and immunoinformatic-designed vaccines targeting VP1, VP2, and conserved non-structural proteins (e.g., 2C) show favorable in silico properties, especially against HRV-C.
- High-valent inactivated or VLP-based formulations are being tested preclinically.
- No licensed HRV vaccine; development has been hindered by:
- Risk-group–oriented prevention:
- For high-risk children (premature infants, severe asthma, chronic lung disease) and the elderly:
- Aggressive control of exposures (e.g., daycare outbreaks, hospital cross-transmission).
- Optimization of baseline asthma/COPD therapy to reduce exacerbation risk.
- Emerging omics-based work aims to identify infants with high-risk endotypes of HRV bronchiolitis for tailored prevention strategies.
- For high-risk children (premature infants, severe asthma, chronic lung disease) and the elderly:
Conclusion
Human rhinoviruses are small, non-enveloped, positive-sense RNA viruses of the Enterovirus genus that cause most common colds and a substantial proportion of lower respiratory tract disease, especially in children and high-risk adults. Their simple structural organization belies extensive antigenic diversity and complex interactions with host immunity, including potent interferon responses, asthma pathogenesis, and viral interference with other respiratory pathogens. Diagnosis is now dominated by sensitive RT-PCR and multiplex platforms, but distinguishing clinically relevant infection from asymptomatic carriage can be challenging. No specific antivirals or vaccines are yet available, although multiple capsids, protease, polymerase, host-targeted, and RNA-targeted strategies are under active investigation, and multi-epitope vaccine designs, particularly for HRV-C, are emerging. Effective control in the near term will continue to rely on supportive care, optimized management of chronic lung diseases, and basic infection-prevention measures, while ongoing advances in structural virology, immunology, and vaccinology offer credible prospects for targeted HRV therapies and vaccines in the longer term.
References
- Rahajamanana, V. L., Thériault, M., Rabezanahary, H., Sahnoun, Y. G., Mallet, M. C., Isabel, S., Trottier, S., & Baz, M. (2025). Advances in the treatment of Enterovirus-D68 and rhinovirus respiratory infections. Infectious Disease Reports, 17(3), 401–415.
- Valbuena, A., Strobl, K., Gil-Redondo, J. C., Valiente, L., de Pablo, P. J., & Mateu, M. G. (2023). Single-molecule analysis of genome uncoating from individual human rhinovirus particles, and modulation by antiviral drugs. Small, 19(44), 2303155.
- Ljubin-Sternak, S., Meštrović, T., Lukšić, I., Mijač, M., & Vraneš, J. (2021). Seasonal coronaviruses and other neglected respiratory viruses: A global perspective and a local snapshot. Frontiers in Public Health, 9, 660597.
- Lee, E., & Gern, J. E. (2025). Clinical significance of rhinoviruses and progress toward vaccination. Allergy, Asthma & Immunology Research, 17(4), 345–361.
- Kiseleva, I., & Ksenafontov, A. (2021). COVID-19 shuts doors to flu but keeps them open to rhinoviruses. Biology, 10(8), 738.
- Zhang, N., Wang, L., Deng, X., Liang, R., Su, M., He, C., Hu, L., Su, Y., Ren, J., Yu, F., Du, L., & Jiang, S. (2020). Recent advances in the detection of respiratory virus infection in humans. Journal of Medical Virology, 92(4), 408–417.
- Maitra, S., Rajak, J., Ghoshal, A., Roy, B., Ghosh, S., Mitra, A., Kumer, A., & Dhara, B. (2025). Rhinovirus, an age-old problem yet to be solved: A comprehensive review discussing modern therapeutics. Health Science Reports, 8(3), e2001.
- Bosch, A., Carcereny, A., García-Pedemonte, D., Fuentes, C., Costafreda, M. I., Pintó, R. M., & Guix, S. (2024). Human enteroviruses and the long road to acute flaccid paralysis eradication. Journal of Applied Microbiology, 136(6), 3159–3177.
- Papaneophytou, C. (2024). Breaking the chain: Protease inhibitors as game changers in respiratory virus management. International Journal of Molecular Sciences, 25(15), 8392.
- Singh, S., Mane, S., Kasniya, G., Cartaya, S., Rahman, M. M., Maheshwari, A., Motta, M., & Dudeja, P. (2022). Enteroviral infections in infants. Newborn, 1(3), 246–261.
- Bizot, E., Bousquet, A., Charpié, M., Coquelin, F., Lefevre, S., Le Lorier, J., Patin, M., Sée, P., Sarfati, E., Walle, S., Visseaux, B., & Basmaci, R. (2021). Rhinovirus: A narrative review on its genetic characteristics, pediatric clinical presentations, and pathogenesis. Frontiers in Pediatrics, 9, 643219.
- Wang, J., Hu, Y., & Zheng, M. (2021). Enterovirus A71 antivirals: Past, present, and future. Acta Pharmaceutica Sinica B, 11(8), 2370–2394.
- Real-Hohn, A., & Blaas, D. (2021). Rhinovirus inhibitors: Including a new target, the viral RNA. Viruses, 13(9), 1835.
- Mamun, T. I., Ali, M. A., Hosen, M. N., Rahman, J., Islam, M. A., Akib, M. G., Zaman, K., Rahman, M. M., Hossain, F., Ibenmoussa, S., Bourhia, M., & Dawoud, T. M. (2025). Designing a multi-epitope vaccine candidate against human rhinovirus C utilizing an immunoinformatics approach. Frontiers in Immunology, 16, 1345678.
- Kumar, N., Fazal, A. D., Panja, S. K., & Kumar, S. (2025). Azithromycin: Molecular and clinical insights into its antiviral and immunomodulatory actions against global health threats. Oriental Journal of Physical Sciences, 10(2), 55–72.
- Ljubin-Sternak, S., & Meštrović, T. (2023). Rhinovirus—A true respiratory threat or a common inconvenience of childhood? Viruses, 15(4), 851.
- Stobart, C. C., Nosek, J. M., & Moore, M. L. (2017). Rhinovirus biology, antigenic diversity, and advancements in the design of a human rhinovirus vaccine. Frontiers in Microbiology, 8, 2412. https://doi.org/10.3389/fmicb.2017.02412
- Khan, M. U., Farman, A., Rehman, A. U., Israr, N., Ali, M. Z. H., & Gulshan, Z. A. (2021). Automated System Design for Classification of Chronic Lung Viruses using Non-Linear Dynamic System Features and K-Nearest Neighbour. Mohammad Ali Jinnah University International Conference on Computing (MAJICC), 1–8. https://doi.org/10.1109/majicc53071.2021.9526272
- Real-Hohn, A., & Blaas, D. (2021). Rhinovirus inhibitors: including a new target, the viral RNA. Viruses, 13(9), 1784. https://doi.org/10.3390/v13091784
