- Influenza A virus (IAV) is an enveloped, negative-sense, segmented RNA virus causing seasonal epidemics and pandemics in humans, with substantial morbidity and mortality worldwide.

- Annual epidemics result in ~1 billion infections and 290,000–650,000 deaths globally.
- IAV also infects birds, swine, horses, dogs, bats and other mammals, with wild aquatic birds serving as the main natural reservoir.
- Key biological features include:
- Eight-segment genome encoding at least 17–18 proteins.
- Rapid evolution via antigenic drift and antigenic shift (reassortment), enabling immune evasion, host switching and drug resistance.
- Understanding structure, replication, pathogenesis, immunity and control measures is critical for postgraduate-level work in virology and public health.
Taxonomy and Classification
- Family: Orthomyxoviridae
- Genus: Alphainfluenzavirus (Influenza A virus).
- Types in family: Influenza A, B, C, D; A and B cause most human disease, A is the only type causing pandemics.
Classification features:
- Subtypes based on HA and NA:
- 18 HA (H1–H18) and 11 NA (N1–N11) glycoprotein subtypes described.
- H1–H16 and N1–N9 circulate in aquatic birds; H17N10 and H18N11 are bat-associated IAV-like viruses.
- Currently endemic in humans: H1N1 and H3N2 seasonal lineages.
- Pathogenicity categories in birds:
- Low pathogenic avian influenza (LPAI): usually mild intestinal/respiratory disease.
- Highly pathogenic avian influenza (HPAI): systemic, often fatal (e.g., H5N1, H5N8, some H7).
- Lineages and clades defined for surveillance (e.g., H5N1 clade 2.3.4.4b) based on HA phylogeny and geographic evolution.
Structure and Morphology
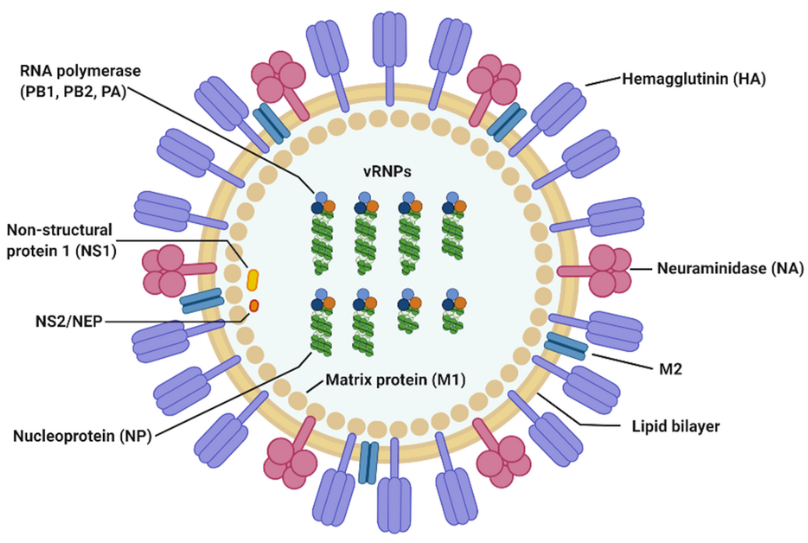
Figure:- Structure and Morphology of Influenza A virus (Source: https://en.wikipedia.org/wiki/Influenza_A_virus)
- Virion properties
- Enveloped, pleomorphic; 80–120 nm in diameter, often spherical in lab strains, filamentous in clinical isolates.
- Lipid bilayer derived from host plasma membrane, studded with HA, NA and fewer M2 channels.
- Surface glycoproteins
- Hemagglutinin (HA):
- Trimeric type I glycoprotein, ~75% of surface spikes.
- Mediates sialic-acid binding and membrane fusion in endosomes.
- Major target of neutralizing antibodies and key determinant of host range and virulence.
- Neuraminidase (NA):
- Tetrameric glycoprotein; cleaves sialic acids to release budding virions and prevent self-aggregation.
- Balances HA binding activity; HA–NA “functional balance” is critical for transmissibility and pathogenicity.
- M2 ion channel:
- Homotetrameric proton channel; low copy number (~1 M2 per 10² HA).
- Acidifies virion interior after endocytosis to dissociate matrix protein from vRNPs.
- Hemagglutinin (HA):
- Internal components
- M1 matrix protein:
- Lines inner envelope; organizes virion structure and binds both membrane and vRNPs.
- Viral ribonucleoproteins (vRNPs):
- Eight helical RNP segments, each containing:
- One negative-sense RNA segment.
- Multiple nucleoprotein (NP) molecules.
- One heterotrimeric polymerase complex: PB2–PB1–PA.
- Eight helical RNP segments, each containing:
- M1 matrix protein:
Genome Organization and Proteins
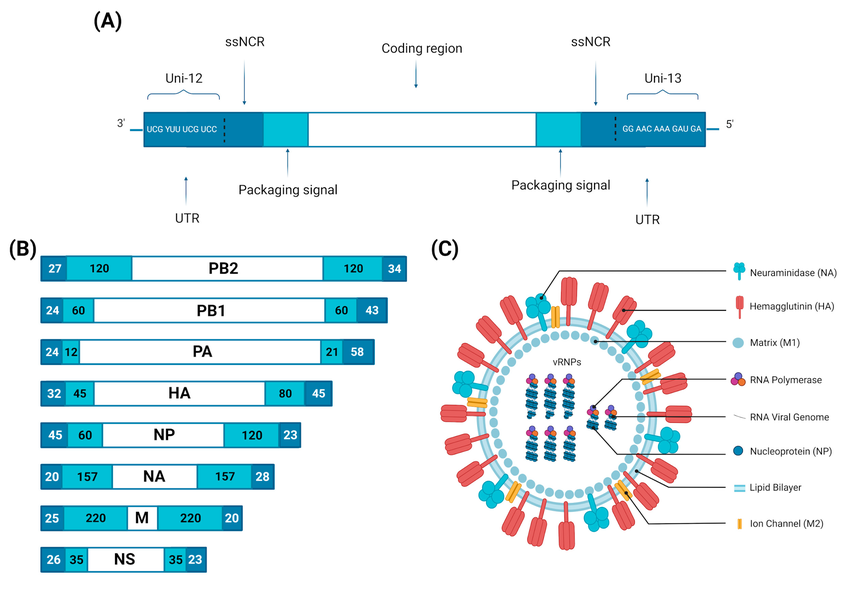
Figure: Influenza A virus virion structure and genome organization. (A) Schematic organization of a gene segment (vRNA) (10.3390/v15040833)
- Genome features
- ~13.5 kb negative-sense, single-stranded RNA, divided into eight segments.
- Each segment packaged into a separate vRNP; transcription and replication occur in the nucleus, unusual for RNA viruses.
- Coding capacity
- At least 17–18 viral proteins generated by alternative splicing, alternative initiation and frameshifting.
Core segments and proteins:
- Segment 1 – PB2
- Polymerase basic 2; cap-binding subunit for “cap-snatching” from host pre-mRNAs.
- Adaptive mutations (e.g., E627K, D701N) enhance replication in mammalian hosts and virulence.
- Segment 2 – PB1
- Catalytic RNA-dependent RNA polymerase; elongation and polyadenylation of viral mRNAs.
- Encodes PB1-F2 (alt ORF):
- Pro-apoptotic mitochondrial protein, contributes to inflammation and virulence.
- Segment 3 – PA
- Endonuclease subunit mediating cleavage of host capped RNAs for priming viral mRNA synthesis.
- Encodes PA-X via ribosomal frameshift:
- Host mRNA shutoff protein, dampens innate and adaptive immunity, modulating virulence.
- Segment 4 – HA
- HA0 precursor cleaved into HA1 (receptor binding) and HA2 (fusion) subunits.
- Cleavage site sequence determines systemic spread:
- Poly-basic cleavage sites in H5/H7 are key determinants of HPAI pathogenesis.
- Segment 5 – NP
- Binds vRNA, forms helical RNP; contains nuclear localization signals for import of vRNPs.
- Also involved in host range and innate immune sensing (e.g., by RIG-I).
- Segment 6 – NA
- Cleaves α-2,3 or α-2,6 sialic acids; mutations can modulate drug resistance and virulence.
- Segment 7 – M
- M1: matrix, essential for assembly, budding, and vRNP nuclear export.
- M2: proton channel critical for uncoating and pH regulation in Golgi during HA maturation.
- Segment 8 – NS
- NS1:
- Multifunctional innate immune antagonist; blocks RIG-I, TLR3, IFN induction and signaling, suppresses host mRNA processing.
- NS2/NEP (nuclear export protein):
- Bridges M1 and cellular export machinery (CRM1), mediating vRNP nuclear export.
- NS1:
Replication Cycle
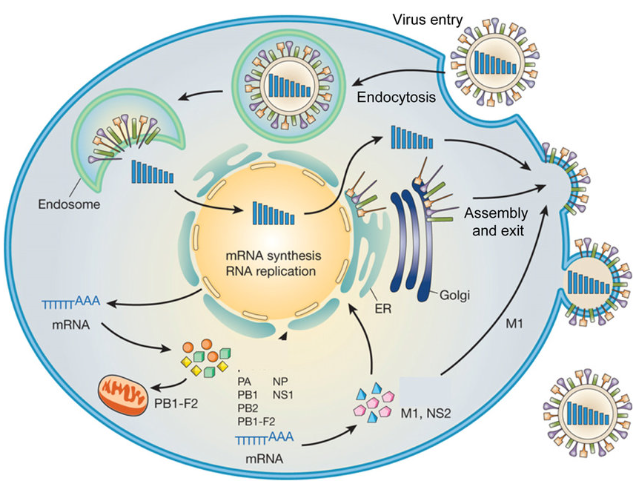
Figure: – Replication Cycle of Influenza A virus (DOI: 10.1016/j.arcmed.2009.10.007)
Overview based largely on:
Attachment and Entry
- Receptor binding
- HA binds terminal sialic acid on host glycoproteins/lipids:
- Human-adapted strains prefer α-2,6-linked sialic acids abundant in upper respiratory tract.
- Avian strains prefer α-2,3 linkages in avian gut and human lower airways.
- HA binds terminal sialic acid on host glycoproteins/lipids:
- Endocytosis
- Virus internalized mainly via clathrin-mediated endocytosis, with contributions from clathrin-independent pathways depending on cell type .
Fusion and Uncoating
- Endosomal acidification
- Low pH triggers:
- Conformational change in HA to fusion-active form; HA2 inserts fusion peptide into endosomal membrane.
- Activation of M2 proton channel, acidifying virion core and dissociating M1 from vRNPs.
- Low pH triggers:
- Fusion
- HA-mediated fusion of viral envelope with endosomal membrane releases vRNPs into cytoplasm.
Nuclear Import and Transcription
- vRNP nuclear import
- NP nuclear localization signals bind importin-α/β, transporting vRNPs to the nucleus.
- Cap-snatching and mRNA synthesis
- PB2 binds 5′ capped ends of host pre-mRNAs; PA endonuclease cleaves 10–13 nt downstream, providing capped primers for PB1.
- PB1 synthesizes capped, polyadenylated viral mRNAs using vRNA as template.
Genome Replication
- Switch from transcription to replication
- Accumulation of NP and polymerase subunits favors synthesis of:
- cRNA (complementary RNA) as full-length positive-sense copy.
- Subsequent synthesis of progeny vRNA from cRNA template.
- Accumulation of NP and polymerase subunits favors synthesis of:
- Both cRNA and vRNA are assembled into RNPs; replication involves FluPol oligomerization, with dimer interfaces required for vRNA synthesis.
Nuclear Export, Assembly, and Budding
- vRNP export
- M1 binds vRNPs; NEP/NS2 recruits CRM1 export machinery, exporting vRNPs to cytoplasm.
- Assembly at plasma membrane
- HA, NA and M2 traffic via secretory pathway to apical membrane.
- M1 organizes budding sites; selective packaging of eight distinct segments via segment-specific packaging signals.
- Budding and release
- Virions bud from apical membrane; NA cleaves sialic acids to prevent self-aggregation and facilitate release.
Pathogenesis and Host Immune Response
Determinants of Virulence and Tropism
- Viral determinants:
- HA receptor specificity and cleavage site (e.g., poly-basic H5/H7).
- Polymerase mutations (PB2 E627K/D701N, PB1, PA changes) enhance replication in mammals.
- NS1, PA-X, PB1-F2 variants modulating host shutoff and inflammation.
- Balance between HA binding and NA cleavage.
- Host determinants
- Age, comorbidities, pregnancy, genetic polymorphisms, and pre-existing immunity critically shape disease severity.
Innate Immunity
- Recognition
- Viral RNAs act as PAMPs sensed by:
- RIG-I, MDA5 in cytoplasm;
- TLR3, TLR7/8 in endosomes;
leading to type I/III IFN and cytokine production.
- Viral RNAs act as PAMPs sensed by:
- Effectors
- IFN-stimulated genes (ISGs) such as MxA, OAS/RNase L, PKR, IFITM proteins restrict viral entry, translation, and replication.
- Recent work identifies host restriction factors targeting polymerase (e.g., DCAF7–CRL4B promoting PA degradation).
- Immune evasion
- NS1 blocks RIG-I signaling, interferes with TRIM25, CPSF30, and polyadenylation, inhibiting IFN and host mRNA maturation.
- Viral manipulation of lncRNAs and circRNAs modulates IFN pathways and cytokine responses.
- IAV can hijack lncRNA IPAN to protect PB1 from RIG-I/TRIM25-mediated degradation, promoting replication.
Adaptive Immunity
- Humoral immunity
- Neutralizing antibodies mainly target HA head; broader antibodies recognize HA stem, NA, and internal antigens.
- Antibody effector functions: neutralization, complement activation, ADCC and ADCP.
- Cell-mediated immunity
- CD8⁺ T cells target conserved epitopes in NP, M1, polymerase; crucial for viral clearance and cross-protection.
- CD4⁺ T cells support B-cell responses and cytotoxic T-cell responses.
- Immunopathology
- Severe disease often involves excessive cytokine responses and damage to respiratory epithelial and endothelial barriers, leading to ARDS and multi-organ dysfunction.
Epidemiology and Transmission
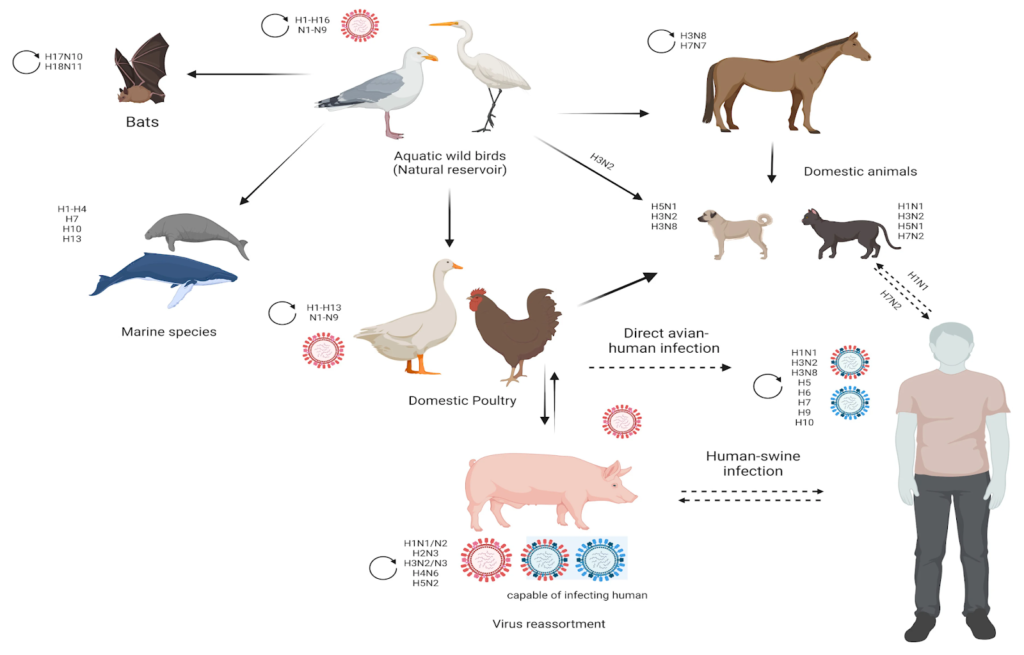
Figure: Schematic diagram showing the host range of influenza A viruses (10.3390/v15040833)
- Global burden
- Seasonal epidemics: 3–5 million severe cases, 290,000–650,000 deaths annually.
- Greatest mortality in elderly, infants, pregnant women, and those with chronic diseases.
- Reservoirs and hosts
- Wild aquatic birds: principal natural reservoir; shedding into water supports fecal–oral transmission in birds.
- Spillover to poultry, swine, horses, dogs, cats, seals, bats and humans, creating complex animal–human interfaces.
- Transmission modes
- In humans: respiratory droplets, aerosols, and contact with contaminated fomites, with virus replicating in respiratory epithelium.
- In birds: mainly fecal–oral, with intestinal replication.
- Evolution and emergence
- Antigenic drift: accumulation of point mutations in HA/NA under immune pressure.
- Antigenic shift (reassortment):
- Co-infection with different IAV strains in a permissive host (e.g., pigs as “mixing vessels”) yields novel subtypes.
- At least five pandemics in the last century (including 1918 H1N1, 1957 H2N2, 1968 H3N2, 2009 H1N1) resulted from reassortment.
Clinical Manifestations
(Primarily based on.)
- Incubation period
- Typically 1–4 days (average 2 days).
- Uncomplicated influenza
- Abrupt onset with:
- Fever, chills, myalgias, headache.
- Nonproductive cough, sore throat, rhinorrhea.
- Fatigue, malaise.
- Illness is usually self-limited (3–7 days), but cough and fatigue can persist for weeks.
- Abrupt onset with:
- Complicated/severe disease
- Viral pneumonia:
- Progressive dyspnea, hypoxemia, bilateral infiltrates; may progress to ARDS.
- Secondary bacterial pneumonia:
- Classically due to Staphylococcus aureus, Streptococcus pneumoniae, or Haemophilus influenzae.
- Biphasic illness: initial improvement then recurrent fever, sputum, respiratory decline.
- Exacerbation of underlying diseases:
- COPD, asthma, heart failure, coronary artery disease, diabetes.
- Viral pneumonia:
- Extra-pulmonary manifestations
- Myositis, rhabdomyolysis, myocarditis, pericarditis.
- Neurologic: febrile seizures, encephalopathy, encephalitis, Guillain–Barré syndrome (rare).
- High-risk groups
- Young children, older adults (≥65 years), pregnant/postpartum women, immunocompromised patients, persons with chronic cardiopulmonary, renal, hepatic, metabolic or neurologic disease, and obesity.
- Avian and zoonotic IAV
- H5N1, H7N9 and related viruses:
- Often cause severe pneumonia, ARDS, multi-organ failure, with high case-fatality rates.
- H5N1, H7N9 and related viruses:
Laboratory Diagnosis
- Specimen types
- Upper respiratory: nasopharyngeal swab/aspirate, throat swab.
- Lower respiratory (severe disease): endotracheal aspirate, BAL.
- Molecular tests
- RT-PCR:
- Gold standard; high sensitivity/specificity, can subtype (H1N1, H3N2, H5, H7, etc.) and differentiate A vs B.
- Multiplex respiratory panels detect influenza plus other viruses.
- RT-PCR:
- Antigen detection
- Rapid antigen tests (RIDTs):
- Provide quick results but lower sensitivity; negative results must be interpreted cautiously in high-prevalence settings.
- Rapid antigen tests (RIDTs):
- Virus isolation
- Cell culture in MDCK or other permissive cells; used for surveillance, antiviral susceptibility testing, and vaccine strain selection.
- Serology
- Hemagglutination-inhibition (HI) and microneutralization tests:
- Mainly for epidemiologic studies, vaccine evaluation, not acute diagnosis.
- Hemagglutination-inhibition (HI) and microneutralization tests:
Treatment and Antiviral Therapy
Currently Approved Antivirals
- Neuraminidase inhibitors (NAIs)
- Oseltamivir (oral), zanamivir (inhaled), peramivir (IV):
- Block NA enzymatic activity → inhibit release of progeny virions.
- Most effective when started within 48 hours of symptom onset; still beneficial later in severe/hospitalized patients.
- Resistance can emerge via NA mutations (e.g., H275Y in N1).
- Oseltamivir (oral), zanamivir (inhaled), peramivir (IV):
- Cap-dependent endonuclease inhibitor
- Baloxavir marboxil:
- Targets PA endonuclease, blocking cap-snatching and viral mRNA synthesis.
- Single-dose oral drug; resistance-associated PA I38X mutations observed after treatment.
- Baloxavir marboxil:
- M2 ion channel inhibitors (adamantanes)
- Amantadine, rimantadine:
- Block M2 proton channel; now largely obsolete due to widespread resistance in circulating IAV strains.
- Amantadine, rimantadine:
Clinical Use
- Indications
- Early treatment of high-risk outpatients.
- All hospitalized or severe cases regardless of onset time.
- Post-exposure prophylaxis in high-risk contacts or institutional outbreaks.
- Combination and novel approaches
- Studies of combination therapy (e.g., NAI + baloxavir) and host-targeted agents (e.g., modulating host proteases, polymerase regulatory factors like DCAF7–CRL4B) are ongoing.
Prevention and Control
- Vaccination – cornerstone of prevention
- Seasonal inactivated or live-attenuated vaccines formulated annually, typically:
- Trivalent or quadrivalent (two IAV subtypes + one or two IBV lineages).
- Vaccine composition updated based on global surveillance of antigenic and genetic drift.
- Efficacy varies by season, age, immune status and match to circulating strains.
- Seasonal inactivated or live-attenuated vaccines formulated annually, typically:
- Limitations
- Antigenic drift and egg-adaptation can reduce vaccine match and effectiveness.
- Production timelines limit rapid response to new pandemic strains.
- Universal vaccine strategies
- Target conserved HA stem, NA, M2e, and internal proteins (NP, M1, polymerase) using:
- Viral vectors, mRNA, nanoparticle and chimeric HA platforms.
- Aim for broad, long-lasting protection across subtypes; several candidates in clinical trials.
- Target conserved HA stem, NA, M2e, and internal proteins (NP, M1, polymerase) using:
- Non-pharmaceutical interventions
- Respiratory hygiene, hand washing, masks, ventilation, staying home when ill.
- Infection control in healthcare and long-term care facilities.
- Culling and biosecurity measures in poultry to control HPAI outbreaks.
- Surveillance and pandemic preparedness
- Global networks (e.g., WHO GISRS) perform virologic and epidemiologic surveillance, antiviral resistance monitoring, and risk assessment of zoonotic strains.
- Stockpiling of antivirals and vaccines, and development of rapid manufacturing platforms are key to pandemic planning.
Conclusion
Influenza A virus is a highly adaptable zoonotic pathogen whose segmented genome, rapid evolution, and broad host range underpin its capacity to cause annual epidemics and periodic, sometimes catastrophic, pandemics. The virus’s structure—particularly HA, NA, M2, and the PB2–PB1–PA polymerase complex—determines host range, transmissibility, and virulence, while accessory proteins such as NS1, PA-X and PB1-F2 fine-tune immune evasion and pathogenicity. Infection outcomes in humans range from mild upper respiratory illness to severe viral pneumonia, ARDS and systemic complications, shaped by viral genetics, host factors and immune responses. Although neuraminidase inhibitors and baloxavir provide effective antiviral options and seasonal vaccination remains the primary preventive tool, antigenic drift, reassortment and drug resistance continue to challenge control efforts. Advancing structural, molecular and immunologic understanding of IAV, together with improvements in universal vaccines, host-targeted therapies and global surveillance, will be central to mitigating the impact of both seasonal influenza and future pandemics.
References
- AbuBakar, U., Amrani, L., Kamarulzaman, F. A., Karsani, S. A., Hassandarvish, P., & Khairat, J. E. (2023). Avian influenza virus tropism in humans. Viruses, 15(4), 833. https://doi.org/10.3390/v15040833
- Liang, Y. (2023). Pathogenicity and virulence of influenza. Virulence, 14(1), 2223057. https://doi.org/10.1080/21505594.2023.2223057
- Carter, T., & Iqbal, M. (2024). The Influenza a Virus Replication Cycle: A Comprehensive review. Viruses, 16(2), 316. https://doi.org/10.3390/v16020316
- Hsu, A. C. (2018). Influenza Virus: A master tactician in innate immune evasion and novel therapeutic Interventions. Frontiers in Immunology, 9, 743. https://doi.org/10.3389/fimmu.2018.00743
- Chauhan, R. P., & Gordon, M. L. (2022). An overview of influenza A virus genes, protein functions, and replication cycle highlighting important updates. Virus Genes, 58(4), 255–269. https://doi.org/10.1007/s11262-022-01904-w
- Su, G., Chen, Y., Li, X., & Shao, J. (2024). Virus versus host: influenza A virus circumvents the immune responses. Frontiers in Microbiology, 15, 1394510. https://doi.org/10.3389/fmicb.2024.1394510
- Dunaeva, M. N. (2024). Molecular genetic mechanisms of influenza A virus replication and mechanisms of action of etiotropic medications. Antibiot Khimioter = Antibiotics and Chemotherapy, 69(3–4), 73–94. https://doi.org/10.37489/0235-2990-2024-69-3-4-73-94
- Jain, R., Sharma, H., Pena, L., Jit, S., Rathi, B., De Oliveira, R. N., & Verma, M. (2025). Influenza virus: Genomic insights, evolution, and its clinical presentation. Microbial Pathogenesis, 205, 107671. https://doi.org/10.1016/j.micpath.2025.107671
- Neyazi, G. R., Khaidarov, S., Kakimova, A. B., Mukhlis, H., & Mohammadi, M. H. (2024). Pathogenesis and immune evasion Strategies of Influenza A virus: Challenges for therapeutic innovation. European Journal of Theoretical and Applied Sciences, 2(5), 783–790. https://doi.org/10.59324/ejtas.2024.2(5).69
- Yu, L., Jiang, Y., Rang, H., Wang, X., Cai, Y., Yan, H., Wu, S., & Lan, K. (2025). Restriction of influenza A virus replication by host DCAF7-CRL4B axis. Journal of Virology, 99(4), e0013325. https://doi.org/10.1128/jvi.00133-25
- Tripathi, G., Faiyaz, M., Hasan, Z., Khanam, A., & Husain, A. (2020). Gene editing and gene therapies in cancer treatment. In Advances in medical diagnosis, treatment, and care (AMDTC) book series (pp. 205–224). https://doi.org/10.4018/978-1-7998-6530-8.ch005
- Javanian, M., Barary, M., Ghebrehewet, S., Koppolu, V., Vasigala, V., & Ebrahimpour, S. (2021). A brief review of influenza virus infection. Journal of Medical Virology, 93(8), 4638–4646. https://doi.org/10.1002/jmv.26990
- Dou, D., Revol, R., Östbye, H., Wang, H., & Daniels, R. (2018). Influenza A virus cell entry, replication, virion assembly and movement. Frontiers in Immunology, 9, 1581. https://doi.org/10.3389/fimmu.2018.01581
- Dangi, T., & Jain, A. (2012). Influenza Virus: A Brief Overview. Proceedings of the National Academy of Sciences India Section B Biological Sciences, 82(1), 111–121. https://doi.org/10.1007/s40011-011-0009-6
- Chen, X., Liu, S., Goraya, M. U., Maarouf, M., Huang, S., & Chen, J. (2018). Host immune response to influenza a virus infection. Frontiers in Immunology, 9. https://doi.org/10.3389/fimmu.2018.00320
- Smyk, J. M., Szydłowska, N., Szulc, W., & Majewska, A. (2022). Evolution of Influenza Viruses—Drug Resistance, treatment options, and Prospects. International Journal of Molecular Sciences, 23(20), 12244. https://doi.org/10.3390/ijms232012244
- Ju, Y., Li, Y., Zhou, C., & Yu, X. (2025). The role of long non-coding RNAs and circular RNAs in immune evasion of influenza A virus: recent advances. Frontiers in Cellular and Infection Microbiology, 15, 1708828. https://doi.org/10.3389/fcimb.2025.1708828
- Krammer, F., Smith, G. J. D., Fouchier, R. a. M., Peiris, M., Kedzierska, K., Doherty, P. C., Palese, P., Shaw, M. L., Treanor, J., Webster, R. G., & García-Sastre, A. (2018). Influenza. Nature Reviews Disease Primers, 4(1), 3. https://doi.org/10.1038/s41572-018-0002-y
- Fan, H., Walker, A. P., Carrique, L., Keown, J. R., Martin, I. S., Karia, D., Sharps, J., Hengrung, N., Pardon, E., Steyaert, J., Grimes, J. M., & Fodor, E. (2019). Structures of influenza A virus RNA polymerase offer insight into viral genome replication. Nature, 573(7773), 287–290. https://doi.org/10.1038/s41586-019-1530-7
- Paules, C., & Subbarao, K. (2017). Influenza. The Lancet, 390(10095), 697–708. https://doi.org/10.1016/s0140-6736(17)30129-0
- Sun, T., Chen, S., Zhou, R., Guo, S., Ye, Y., Qiu, J., Li, X., Cen, S., & Wang, J. (2025). IncRNA IPAN antagonizes RIG-I/TRIM25-mediated degradation of influenza A virus PB1 to promote viral replication. Biosafety and Health, 7(3), 199–208. https://doi.org/10.1016/j.bsheal.2025.05.005
- Arias, C. F., Escalera-Zamudio, M., De Los Dolores Soto-Del Río, M., Cobián-Güemes, A. G., Isa, P., & López, S. (2009). Molecular Anatomy of 2009 Influenza virus A (H1N1). Archives of Medical Research, 40(8), 643–654. https://doi.org/10.1016/j.arcmed.2009.10.007
