- Influenza B virus (IBV) is a segmented, negative-sense RNA virus that, together with influenza A, causes seasonal influenza in humans.

- IBV lacks known pandemic potential because it primarily infects humans (and a few other mammals) and has no established animal reservoir analogous to wild aquatic birds for influenza A.
- Despite this, IBV can dominate or equal influenza A in some seasons, causing substantial morbidity, particularly in children and adolescents, and can be associated with severe pneumonia and death.
- IBV evolves by antigenic drift and is divided into two major antigenically distinct lineages (B/Victoria and B/Yamagata), complicating vaccine formulation and necessitating quadrivalent vaccines.
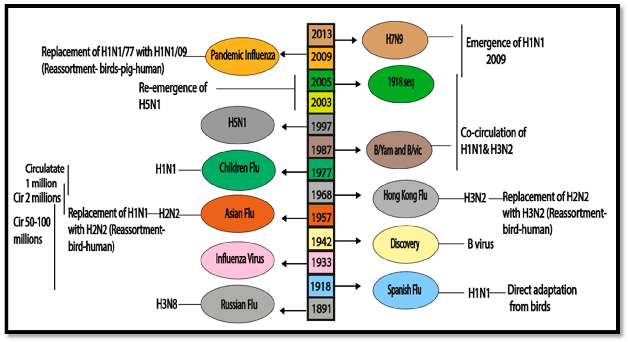
Taxonomy and Classification of Influenza B Virus (IBV)
- Family: Orthomyxoviridae
- Genus: Betainfluenzavirus (Influenza B virus).
- Types in family: Influenza A, B, C, and D; types A and B are the major human respiratory pathogens, with IBV confined largely to humans.
Key classification aspects:
Lineages:
- B/Victoria/2/87-like (Victoria lineage).
- B/Yamagata/16/88-like (Yamagata lineage).
Antigenic and genetic characteristics
- Lineages are based mainly on HA and NA gene phylogeny; each lineage further subdivided into clades/subclades:
- B/Victoria: clade V1A and subclades V1A.1, V1A.2, V1A.3.
- B/Yamagata: clades Y1, Y2, Y3, with no official subclades.
- Victoria lineage shows frequent antigenic drift near the HA receptor-binding site, leading to 1–3-year strain replacement cycles, whereas Yamagata lineage has a more antigenically stable and conservative pattern.
Host range
- Natural host considered humans, but IBV also detected in seals, sea mammals, domesticated and wild dogs, and some birds, without clear evidence of sustained zoonotic cycles back into humans.
Structure and Morphology of Influenza B Virus (IBV)
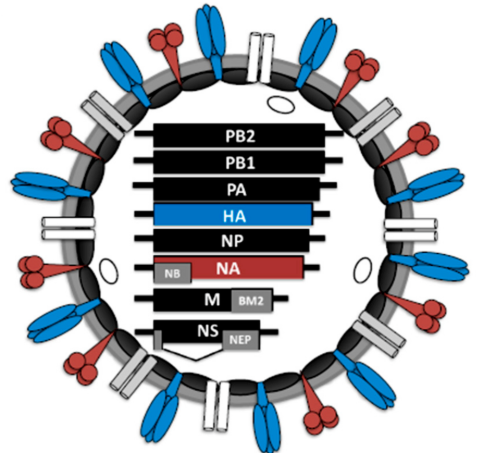
Virion morphology
- Enveloped, roughly spherical or filamentous particles.
- Diameter of spherical virions ~100 nm; filamentous forms may exceed 300 nm, similar to influenza A.
Envelope and matrix
- Lipid envelope derived from host plasma membrane.
- Matrix protein (BM1/M1) forms a layer beneath the envelope, interacting with both glycoproteins and ribonucleoproteins (vRNPs).
Surface glycoproteins
Hemagglutinin (HA):
- Trimeric glycoprotein mediating sialic-acid receptor binding and low-pH-triggered membrane fusion.
- Principal antigenic determinant and main target of neutralizing antibodies.
Neuraminidase (NA):
- NB protein:
- Unique small hydrophobic glycoprotein of IBV, encoded on segment 6; its function is incompletely understood, but it appears to be a membrane protein involved in virion structure or host interaction.
- IBV encodes BM2 as the ion channel, rather than M2 of IAV, and NB replaces the IAV M2 on the surface glycoprotein repertoire.
Internal components
- vRNPs: helical complexes of vRNA with nucleoprotein (NP) and polymerase (PB2, PB1, PA).
- Unlike IAV’s typical “7+1” RNP arrangement, IBV RNPs form more loosely packed bundles without a defined 7+1 pattern.
Genome Organization and Proteins of Influenza B Virus (IBV)
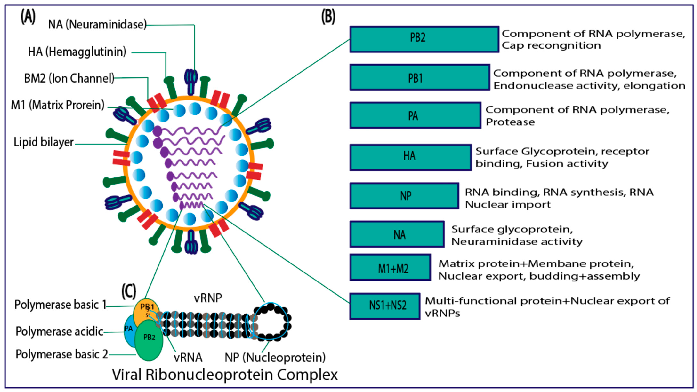
Genome
- Eight segments of negative-sense, single-stranded RNA, total length ~14 kb.
- Each segment is encapsidated with NP and associated with one copy of the heterotrimeric polymerase.
Segment organization and major proteins (IBV generally encodes ~11 proteins, lacking IAV accessory proteins PB1-F2 and PA-X):
Segment 1 – PB2
- ~2300–2400 nt; ~759–760 aa.
- Polymerase basic 2; binds capped host RNAs to initiate transcription (“cap-snatching”).
- Distinct cap-binding characteristics from IAV, with inverted substrate recognition and lower m⁷GDP affinity.
Segment 2 – PB1
- ~2300–2400 nt; ~757–759 aa.
- Catalytic RNA-dependent RNA polymerase, mediates mRNA and cRNA/vRNA synthesis; also possesses endonuclease function together with PA.
Segment 3 – PA
- ~2150–2250 nt; ~716–747 aa.
- Polymerase acidic subunit, involved in protease activity and polymerase assembly.
Segment 4 – HA
- ~1700–1800 nt; 550–583 aa; synthesized as HA0 and cleaved to HA1 (receptor binding) and HA2 (fusion).
- Lineage- and clade-defining mutations cluster in antigenic sites and near the receptor-binding region.
Segment 5 – NP
- ~1500–1600 nt; 498–500 aa.
- Binds vRNA to form RNPs; contains nuclear localization signals; functions in transcription and replication.
Segment 6 – NA and NB
- ~1400–1500 nt.
- Encodes NA (467–470 aa) – sialidase for virion release.
- Overlapping reading frame encodes NB, a small membrane protein with unclear but likely structural or ion-channel-like roles.
Segment 7 – BM1/BM2 (Matrix segment)
- ~1000–1100 nt.
- BM1 (M1): 252–255 aa, matrix protein for virion shape, budding, and vRNP interaction.
- BM2: ion channel, functionally analogous to IAV M2 but structurally and pharmacologically distinct (amantadines are ineffective against IBV BM2).
Segment 8 – NS segment
- ~900–1000 nt.
- NS1: interferon antagonist, modulates host gene expression and innate immune responses.
- NS2/NEP: nuclear export protein regulating vRNP nuclear export.
- IBV does not encode PA-X or PB1-F2, and its accessory protein repertoire is less complex than IAV’s, with differences in length and amino-acid compositions across most proteins.
Replication Cycle of Influenza B Virus (IBV)
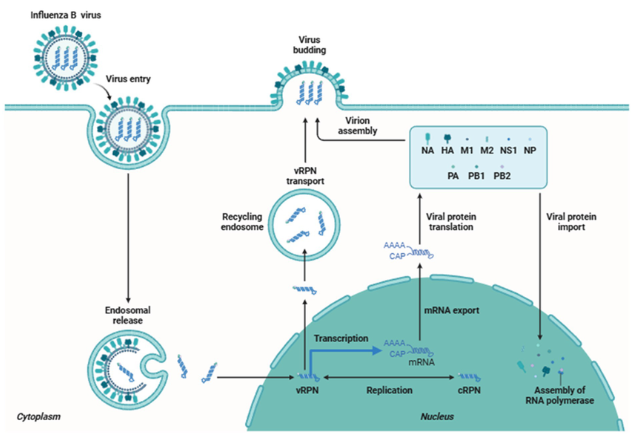
(Parallels IAV but with IBV-specific features).
Attachment and entry
- HA binds terminal sialic acids on host cell surface glycoproteins and glycolipids.
- IBV recognizes primarily α2,6-linked sialic acids in human upper respiratory tract, similar to human-adapted IAV.
- Virus enters via clathrin-mediated endocytosis and other endocytic pathways.
Fusion and uncoating
Endosomal acidification triggers:
- HA conformational change → fusion of viral envelope with endosomal membrane.
- BM2 ion channel activation → proton influx into virion, disassembly of BM1–vRNP interactions, and release of vRNPs.
Nuclear import and transcription
- vRNPs imported into nucleus via NP nuclear localization signals and importings.
- PB2 binds capped host pre-mRNA; PA/PB1 mediate cap-snatching and synthesis of capped, polyadenylated viral mRNAs.
Genome replication
- Early phase favors mRNA transcription; as NP and polymerase accumulate, switch to replication:
- vRNA → full-length cRNA → progeny vRNA, both assembled into RNPs.
- IBV polymerase interacts with host RNA polymerase II and ANP32 cofactors via distinct interfaces compared with IAV, suggesting type-specific differences in RNA synthesis and RNP assembly.
- Specific PB2/NP substitutions (e.g., PB2-N460S and NP-I163T) synergistically enhance polymerase activity, viral replication and pathogenicity in vitro and in vivo.
Nuclear export, assembly and budding
- NS2/NEP mediates CRM1-dependent nuclear export of vRNPs via BM1–vRNP interactions.
- HA, NA, NB and BM2 traffic to plasma membrane; BM1 organizes assembly.
- Eight distinct vRNPs are selectively packaged, though IBV shows different vRNP spatial organization compared with IAV.
- Virions bud from apical surface; NA cleaves sialic acids to release virions.
Pathogenesis and Host Immune Response of Influenza B Virus (IBV)
Viral determinants of pathogenicity
Structural and polymerase determinants
- HA receptor affinity, stability and cleavage properties influence upper vs lower respiratory tract tropism and severity.
- PB2-N460S and NP-I163T substitutions synergistically increase polymerase activity, viral replication and pathogenicity, associated with exaggerated lung inflammation and severe disease in mice.
NS1 and innate immune antagonism
- IBV NS1 is essential for efficient growth and interferon antagonism, inhibiting IFN-β induction and interacting with host factors including ISG15 and nuclear speckles.
- NS1 contributes significantly to evasion of RIG-I-mediated antiviral responses and shaping cytokine profiles.
Tissue tropism and lesions
- IBV primarily infects respiratory epithelium (ciliated, goblet, club, and basal cells) in bronchi and bronchioles.
- In severe seasons (e.g., 2017/18 Yamagata-dominant), IBV strains show improved replication fitness, tolerance to higher temperatures and extended tropism to type I and II alveolar cells and alveolar macrophages, supporting lower-respiratory-tract adaptation.
Histopathology in severe/fatal cases:
- Necrotizing tracheobronchitis, ulceration, sloughing of mucosa.
- Diffuse alveolar damage, hyaline membranes, hemorrhage, and limited neutrophil infiltration.
Innate and adaptive immunity
Innate response
- Rapid induction of type I and III IFNs and pro-inflammatory cytokines; however, some IBV strains attenuate IFN responses via NS1 and polymerase-driven mechanisms.
Adaptive response
Humoral immunity:
- Neutralizing antibodies mainly target HA, with NA also contributing to protective immunity.
T-cell immunity:
- Major CD8⁺ T-cell antigens are less well defined than for IAV; NP and M1 are believed to contribute, but detailed immunodominance hierarchies remain uncertain.
Epidemiology and Transmission
Burden and age distribution
- IBV contributes substantially to seasonal influenza burden, sometimes dominating seasons and causing disease impact similar to influenza A.
- Disproportionately affects children and adolescents, with higher hospitalization and complication rates compared with adults.
Global circulation
- B/Victoria and B/Yamagata co-circulated for decades; recent years show global decline or apparent disappearance of B/Yamagata since 2019, with B/Victoria predominating in multiple regions.
- Molecular surveillance in Africa and Brazil demonstrates alternating lineage predominance and turnover of clades/subclades, especially B/Victoria V1A, V1A.3 and related clades, driven by HA antigenic mutations.
Lineage-specific evolution
B/Victoria:
- Exhibits rapid antigenic drift with monophyletic strain replacement every 1–3 years.
B/Yamagata:
- More antigenically conserved, but associated with severe seasons (e.g., 2017/18 in Europe) when combined with vaccine mismatch and viral adaptation to lower respiratory tract.
Transmission
- Primarily via respiratory droplets, aerosols, and contact with contaminated fomites, similar to influenza A.
- Intranasal route leads to infection of upper airway epithelium, replication, shedding, and person-to-person spread.
Clinical Manifestations
Typical illness
- Sudden onset fever, myalgia, headache, malaise, dry cough, sore throat, rhinorrhea; clinically indistinguishable from influenza A.
Severity spectrum
- Most cases: mild to moderate respiratory disease in otherwise healthy individuals.
B/Yamagata:
- Pneumonia, ARDS, respiratory failure, secondary bacterial infections, exacerbations of asthma/COPD, and, rarely, death, especially in high-risk groups.
Burden in children
- IBV causes a significant proportion of pediatric influenza hospitalizations; young children often experience more severe disease than adults.
- Extra-pulmonary manifestations reported: febrile seizures, encephalopathy, myocarditis and myositis, although less extensively studied than in IAV.
Lineage-specific clinical patterns
In Thai tertiary-care data (2011-2014), Victoria lineage infection was significantly associated with:
Longer hospitalization, higher pneumonia incidence (25% vs 6.1% for Yamagata), more secondary bacterial infection, and more underlying disease among hospitalized cases.
Recent pneumonia data
- In Beijing 2021–2022, 11.7% of influenza B-positive outpatients developed pneumonia; a minority required hospitalization (2.7%) and mortality was low (0.5%), with B/Victoria clade 1A.3 viruses predominating and harboring characteristic HA/NA mutations.
Laboratory Diagnosis
Specimens
- Nasopharyngeal swabs/aspirates, throat swabs; in severe disease, lower respiratory tract samples (e.g., BAL).
Molecular detection
RT-PCR is the diagnostic gold standard:
- Pan-influenza A/B assays, followed by lineage-specific or HA/NA sequencing for B/Victoria vs B/Yamagata assignment.
- Universal IBV genomic assays (segment-specific RT-PCR) enable whole-genome sequencing and reverse genetics.
Virus isolation
- Culture in MDCK or other permissive cells for antigenic characterization, antiviral susceptibility testing, and vaccine strain selection.
Serology
- Hemagglutination-inhibition (HI) and microneutralization assays used in vaccine evaluation and sero-epidemiology, not routine diagnosis.
Treatment and Antiviral Therapy
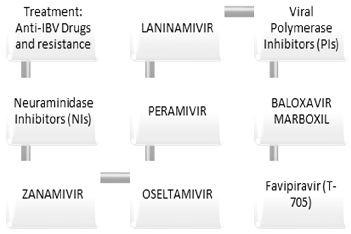
Burden and therapeutic need
- IBV causes substantial morbidity and mortality, particularly in children, yet clinical awareness and study of treatment efficacy lag behind influenza A.
Neuraminidase inhibitors (NAIs)
Oseltamivir, zanamivir, peramivir:
- Inhibit NA, blocking viral release; clinically effective against IBV, reducing duration and severity when started early.
- Resistance has been documented, sometimes without clear fitness cost, but prevalence remains generally low.
- Clinical data for IBV were historically weaker than for IAV, leading to uncertainty; more recent studies and meta-analyses support the benefit, especially in children and high-risk patients.
Polymerase inhibitor (baloxavir marboxil)
- Cap-dependent endonuclease inhibitor targeting PA; effective against IBV.
- Large randomized trials in high-risk influenza B patients show significant reductions in viral load and symptom duration; baloxavir may be more effective than oseltamivir for IBV in some analyses.
Adamantanes
- Amantadine and rimantadine target the M2 channel of IAV and are ineffective against IBV BM2; not recommended for IBV.
Resistance and future options
- IBV’s high mutation and replication rates facilitate emergence of resistance to NAIs and baloxavir, necessitating ongoing surveillance.
- Novel antivirals and host-targeted strategies (e.g., polymerase-host factor interactions, BM2 targeting) are under exploration.
Prevention and Control
Vaccination
Seasonal influenza vaccines (inactivated, live-attenuated) include IBV antigens:
- Trivalent: one IBV lineage.
- Quadrivalent: both B/Victoria and B/Yamagata, now standard because mismatches between vaccine lineage and circulating IBV cause marked loss of protection.
- Overall influenza vaccine effectiveness is ~38%, with ~50% effectiveness against IBV in some analyses, but strongly dependent on antigenic match.
Vaccine mismatch and evolution
- 2017/18 season: Yamagata-dominant IBV with vaccine mismatch contributed to unusually severe disease; intrinsic viral adaptations (lower-tract tropism and immune evasion) also played a role.
- Recent seasons show disappearance or near-absence of B/Yamagata in some settings, raising questions about future quadrivalent vaccine design.
Cross-protective and eradication-oriented vaccines
Cross-protective strategies include:
- Recombinant viruses, VLPs, recombinant HA/NA/NP/MB1 proteins, and conserved epitope-based vaccines.
- Targets: conserved regions of HA (stem), NA, NP, BM1 and possibly NB, aiming for broad protection across lineages and clades.
Because IBV is essentially restricted to humans, effective cross-protective vaccination could theoretically enable eradication, making IBV a particularly attractive target for next-generation vaccines.
Non-pharmaceutical interventions
- Standard respiratory infection control measures (hand hygiene, masks, respiratory etiquette, isolation of symptomatic individuals) reduce IBV transmission.
Surveillance
- Global and regional genomic surveillance of IBV lineages, clades, antigenic drift and antiviral resistance is crucial for:
- Vaccine strain selection,
- Early identification of high-virulence strains,
- Guiding antiviral use.
Conclusion
Influenza B virus, long overshadowed by influenza A due to its lack of pandemic reassortment potential, is now recognized as a major human respiratory pathogen with a distinctive biology and considerable clinical impact. Its human-adapted ecology, dual-lineage structure, and antigenic drift patterns create recurrent seasons in which IBV, particularly in children and adolescents, contributes substantially to hospitalizations, pneumonia, and, occasionally, death. At the molecular level, IBV shares the overall orthomyxovirus architecture and nuclear replication strategy of IAV but differs in accessory proteins, polymerase interfaces, RNP organization and the presence of NB and BM2, which together shape its replication and pathogenicity. Recent work has begun to link specific polymerase and NP mutations to enhanced replication and virulence, and lower-respiratory-tract adaptation has been documented in severe seasons. Clinically, neuraminidase inhibitors and baloxavir are effective therapeutic options, while quadrivalent vaccines and emerging cross-protective candidates form the backbone of prevention. Because IBV is largely restricted to humans, sustained improvements in vaccination, antiviral stewardship and surveillance may not only reduce seasonal burden but also open the long-term possibility of influenza B eradication.
References
- Din, G. U., Hasham, K., Amjad, M. N., & Hu, Y. (2023). Natural History of Influenza B Virus—Current knowledge on treatment, resistance and therapeutic options. Current Issues in Molecular Biology, 46(1), 183–199. https://doi.org/10.3390/cimb46010014
- Ashraf, M. A., Raza, M. A., Amjad, M. N., Din, G. U., Yue, L., Shen, B., Chen, L., Dong, W., Xu, H., & Hu, Y. (2024). A comprehensive review of influenza B virus, its biological and clinical aspects. Frontiers in Microbiology, 15, 1467029. https://doi.org/10.3389/fmicb.2024.1467029
- Hutchinson, E. C. (2018). Influenza virus. Trends in Microbiology, 26(9), 809–810. https://doi.org/10.1016/j.tim.2018.05.013
- Krammer, F., Smith, G. J. D., Fouchier, R. a. M., Peiris, M., Kedzierska, K., Doherty, P. C., Palese, P., Shaw, M. L., Treanor, J., Webster, R. G., & García-Sastre, A. (2018). Influenza. Nature Reviews Disease Primers, 4(1), 3. https://doi.org/10.1038/s41572-018-0002-y
- Zaraket, H., Hurt, A. C., Clinch, B., Barr, I., & Lee, N. (2020). Burden of influenza B virus infection and considerations for clinical management. Antiviral Research, 185, 104970. https://doi.org/10.1016/j.antiviral.2020.104970
- Koutsakos, M., Nguyen, T. H., Barclay, W. S., & Kedzierska, K. (2015). Knowns and unknowns of Influenza B viruses. Future Microbiology, 11(1), 119–135. https://doi.org/10.2217/fmb.15.120
- Din, G. U., Wu, C., Tariq, Z., Hasham, K., Amjad, M. N., Shen, B., Yue, L., Raza, M. A., Ashraf, M. A., Chen, L., & Hu, Y. (2024). Unlocking influenza B: exploring molecular biology and reverse genetics for epidemic control and vaccine innovation. Virology Journal, 21(1), 196. https://doi.org/10.1186/s12985-024-02433-8
- Bhat, Y. R. (2020). Influenza B infections in children: A review. World Journal of Clinical Pediatrics, 9(3), 44–52. https://doi.org/10.5409/wjcp.v9.i3.44
- Liang, Y. (2023). Pathogenicity and virulence of influenza. Virulence, 14(1), 2223057. https://doi.org/10.1080/21505594.2023.2223057
- Paules, C., & Subbarao, K. (2017). Influenza. The Lancet, 390(10095), 697–708. https://doi.org/10.1016/s0140-6736(17)30129-0
- Nyarango, B. K., Owuor, D. C., Isoe, E., Mutunga, M., Cheruiyot, R., Katama, E. N., Makori, T., Lambisia, A., Nyiro, J., Githinji, G., Kiguli, S., Olupot-Olupot, P., Maitland, K., & Agoti, C. N. (2025). Epidemiological dynamics of influenza B virus across multiple seasons in Kenya and Uganda inferred from sequence data, 2010–2021. BMC Infectious Diseases, 25(1), 1080. https://doi.org/10.1186/s12879-025-11442-z
- Wang, Y., Gao, Y., Ma, T., Ye, Y., Cao, C., Zou, B., Ye, S., Huang, Q., Yang, C., Li, S., Liang, L., Zhou, H., Zeng, Z., Yang, Z., & Pan, W. (2025). N460S in PB2 and I163T in nucleoprotein synergistically enhance the viral replication and pathogenicity of influenza B virus. PLoS Pathogens, 21(9), e1013463. https://doi.org/10.1371/journal.ppat.1013463
- Wang, Y., Liu, Y., Wang, Y., Mai, H., Chen, Y., Zhang, Y., Ji, Y., Cong, X., & Gao, Y. (2024). Clinical characteristics of outpatients with influenza-B-associated pneumonia and molecular evolution of influenza B virus in Beijing, China, during the 2021–2022 influenza season. Archives of Virology, 169(2), 30. https://doi.org/10.1007/s00705-023-05957-6
- Tsybalova, L. M., Stepanova, L. A., Ramsay, E. S., & Vasin, A. V. (2022). Influenza B: Prospects for the development of Cross-Protective Vaccines. Viruses, 14(6), 1323. https://doi.org/10.3390/v14061323
- Brcko, I. C., De Souza, V. C., Ribeiro, G., Lima, A. R. J., Martins, A. J., Barros, C. R. D. S., De Carvalho, E., Pereira, J. S., De Lima, L. P. O., Viala, V. L., Kashima, S., De La Roque, D. G. L., Santos, E. V., Rodrigues, E. S., Nunes, J. A., Torres, L. S., Caldeira, L. a. V., Palmieri, M., Medina, C. G., . . . Elias, M. C. (2024). Comprehensive molecular epidemiology of influenza viruses in Brazil: insights from a nationwide analysis. Virus Evolution, 11(1), veae102. https://doi.org/10.1093/ve/veae102
- Horthongkham, N., Athipanyasilp, N., Pattama, A., Kaewnapan, B., Sornprasert, S., Srisurapanont, S., Kantakamalakul, W., Amaranond, P., & Sutthent, R. (2016). Epidemiological, Clinical and Virological Characteristics of Influenza B Virus from Patients at the Hospital Tertiary Care Units in Bangkok during 2011-2014. PLoS ONE, 11(7), e0158244. https://doi.org/10.1371/journal.pone.0158244
- Çalışkan, D. M., Kumar, S., Hinse, S., Schughart, K., Wiewrodt, R., Fischer, S., Krueger, V., Wiebe, K., Barth, P., Mellmann, A., Ludwig, S., & Brunotte, L. (2024). Molecular characterisation of influenza B virus from the 2017/18 season in primary models of the human lung reveals improved adaptation to the lower respiratory tract. Emerging Microbes & Infections, 13(1), 2402868. https://doi.org/10.1080/22221751.2024.2402868
- Hutchinson, E. C. (2018). Influenza virus. Trends in Microbiology, 26(9), 809–810. https://doi.org/10.1016/j.tim.2018.05.013
