Orthopoxviruses are large DNA viruses of major historical and current public-health importance, exemplified by variola virus (smallpox), vaccinia virus, cowpox virus, and, most recently, mpox (monkeypox) virus. The 2022–2025 global mpox outbreaks have renewed interest in orthopoxvirus biology, pathogenesis, and control.

- Large, enveloped, double-stranded DNA (dsDNA) viruses, 200–250 nm in size.
- Replicate exclusively in the cytoplasm, despite being DNA viruses.
- Encode a large repertoire of their own transcription, replication, and immune-evasion proteins, minimizing dependence on host nuclear functions.
- Include multiple human and zoonotic pathogens with cross-reactive antigenicity, enabling cross-protection by smallpox vaccines against mpox and other orthopoxviruses.
Taxonomy and Classification
- Family: Poxviridae
Subfamily: Chordopoxvirinae
Genus: Orthopoxvirus.
- Medically relevant orthopoxviruses
Variola virus (VARV) – smallpox.
Mpox virus (MPXV; formerly monkeypox virus).
Vaccinia virus (VACV) – vaccine and laboratory strain.
Cowpox virus (CPXV).
Others: camelpox, ectromelia (mousepox).
- Genus-level features
Enveloped, brick- or oval-shaped virions with linear dsDNA genomes of ~186–228 kbp encoding ~174–233 proteins.
Conserved central genomic region: essential “housekeeping” genes for replication, transcription, and virion assembly.
Variable terminal regions: host range, virulence, immunomodulatory proteins.
- Recent MPXV classification (ICTV, WHO)
Clade I (Congo Basin; more virulent).
Clade II (West African) is subdivided into IIa and IIb; IIb B.1 lineage dominates 2022+ outbreaks.
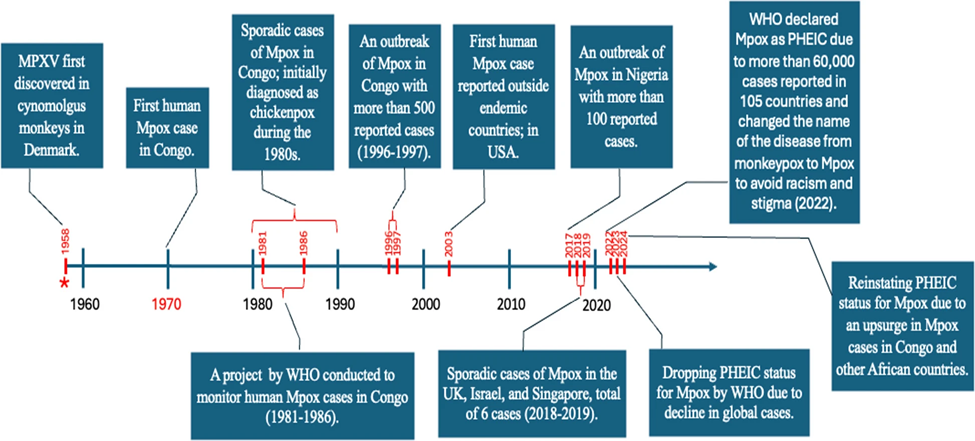
Fig 1: The timeline of Mpox outbreaks and major events (Hajjo et al., 2025)
Structure and Morphology
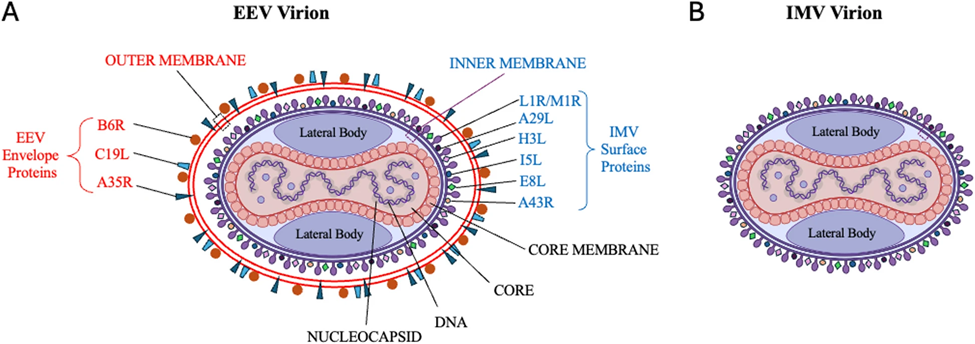
Fig 2: A schematic showing the enveloped viral particle of MPXV and key structural proteins. A) EEV virion. B) IMV virion (Hajjo et al., 2025)
Orthopoxvirions share a characteristic architecture:
- Virion size and shape
- ~200–250 nm in length; brick- or ovoid-shaped particles with a lipoprotein envelope.
- Core and lateral bodies
- Central dumbbell-shaped core containing dsDNA genome and core enzymes.
- Two lateral bodies flanking the core; deliver early enzymes and immune-modulatory factors into the cytoplasm on entry.
- Outer structures
- Core membrane, surrounded by a palisade layer → nucleoprotein complex.
- One or two lipid envelopes depending on virion form (IMV vs EEV).
Key infectious forms (best characterized in MPXV and VACV):
- Intracellular mature virus (IMV)
- Single membrane.
- Environmentally stable; mediates host-to-host and fomite transmission.
- Intracellular enveloped virus (IEV)
- IMV wrapped in additional Golgi-/endosome-derived membranes.
- Cell-associated enveloped virus (CEV)
- IEV that reaches the plasma membrane and remains attached; promotes cell-to-cell spread.
- Extracellular enveloped virus (EEV)
- Released CEV; important for long-range dissemination and immune evasion.
Genome Organization and Proteins
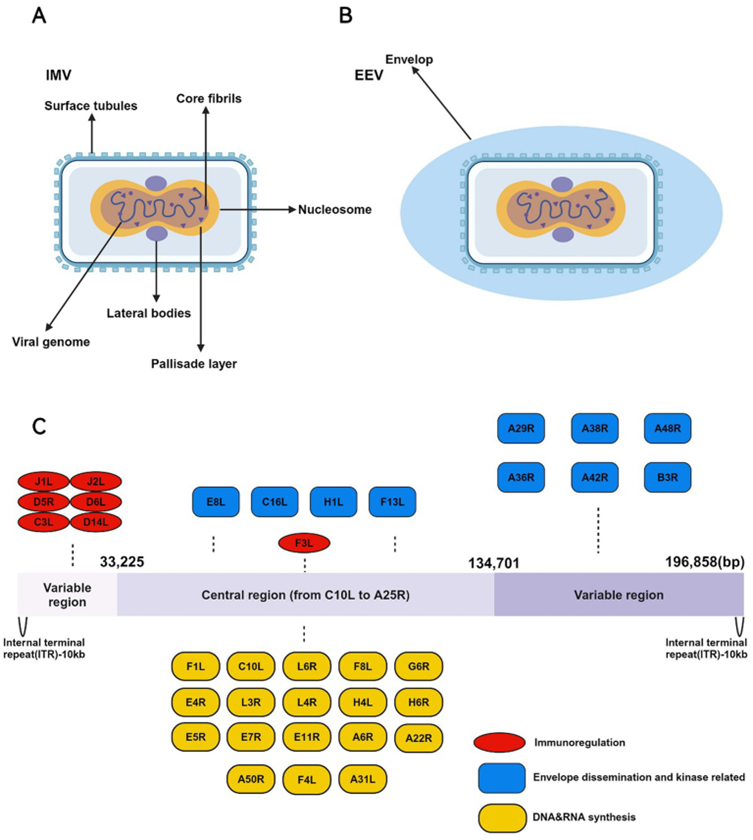
Fig 3: The IMV and EEV morphology and genomic structure of the MPXV (Chen et al., 2025)
- Genome type and size
- Linear double-stranded DNA (dsDNA), ~186–228 kbp; MPXV ≈197 kbp; VACV ≈195 kbp; CPXV up to ≈220 kbp.
- Encodes ~170–230 open reading frames (ORFs), depending on species and strain.
- Terminal hairpins and inverted terminal repeats (ITRs)
- Ends covalently closed as hairpin telomeres; no free 3′ or 5′ ends.
- Each terminus contains an ITR region (≈6–10 kbp) with:
- Tandem repeats and low-complexity sequences.
- Duplicated genes (one copy in each ITR), contributing to gene dosage.
- Hairpins contain loops and mismatched bases; structural integrity (not exact sequence) is essential for maturation and packaging.
- Conserved central vs variable terminal regions
- Central “core” region (~120–130 kbp):
- High conservation of gene order and sequence across orthopoxviruses.
- Encodes essential genes for DNA replication, transcription, and virion morphogenesis.
- Terminal regions:
- Highly variable in content and length; major site of gene gains, losses, duplications, and rearrangements.
- Enriched for host-range factors, virulence genes, and immune modulators.
- Central “core” region (~120–130 kbp):
- Core vs accessory genes (OPG nomenclature)
- About half of the orthopox genes are core/essential (replication, expression, morphogenesis).
- The remaining ≈100 are accessory genes, usually non-essential in vitro but crucial for host interaction and immune evasion.
- Unified Orthopoxvirus Gene (OPG) system numbers genes from OPG001 (left end) to OPG214 (right end).
- Adaptive evolution pattern
- Central replication/morphogenesis genes are largely under purifying selection, but some show strong positive selection (e.g., B1R, A24R, H3L).
- Terminal genes under diversifying selection, especially those encoding immunomodulators and virion-associated host-interaction proteins.
- Intrinsically disordered regions (IDRs) are common and evolve fast; many positively selected sites map to IDRs.
Functional Gene Categories
- Replication and repair (central region)
- E9: DNA-dependent DNA polymerase.
- A20 (A22/A20 complex in MPXV): processivity factor; exapted from inactivated NAD-dependent DNA ligase.
- D4: uracil-DNA glycosylase (E4 in MPXV) with base-excision repair (BER) function.
- D5: primase–helicase/NTPase required for origin unwinding and primer synthesis.
- B1: protein kinase involved in replication/repair regulation.
- Transcription and mRNA processing
- Multi-subunit RNA polymerase complex.
- Early, intermediate, and late transcription factors (e.g., D10, J6).
- Capping, methyltransferase, poly(A) polymerase and termination proteins.
- Structural and morphogenesis genes
- Major core proteins (e.g., A3; exapted deubiquitinase), membrane proteins, scaffold proteins (D13), assembly factors.
- Proteins for crescent/immature virion formation, DNA packaging, and proteolytic maturation of core proteins.
- Host-interaction and immune-evasion genes (terminals)
- Secreted cytokine decoys (TNF, IL-1, IFN binding proteins).
- Chemokine-binding proteins, complement regulator (e.g., D14/MOPICE).
- Ankyrin-repeat and kelch-like proteins modulating host signaling and cell cycle.
- Many under-diversifying or episodic positive selection, especially on MPXV lineages.
- MPXV-specific points
- Genome ≈197 kb, ≈200 predicted genes with conserved central block and variable terminals.
- OPG027 (C7L family) and D1L are strongly implicated in host adaptation, transmissibility, and immune evasion; several 2022-outbreak lineages show mutations here.
- Tandem repeat and LCR variation (e.g., OPG153, OPG204, OPG208) acts as a “genomic accordion” to rapidly adjust host-interaction traits.
Replication Cycle
Despite being dsDNA viruses, orthopoxviruses replicate exclusively in the cytoplasm, where they form “viral factories.”
- Entire replication occurs in the cytoplasm within discrete foci called viral factories/virosomes.
- Orthopoxviruses encode virtually all proteins required for DNA replication, repair, recombination, and expression.
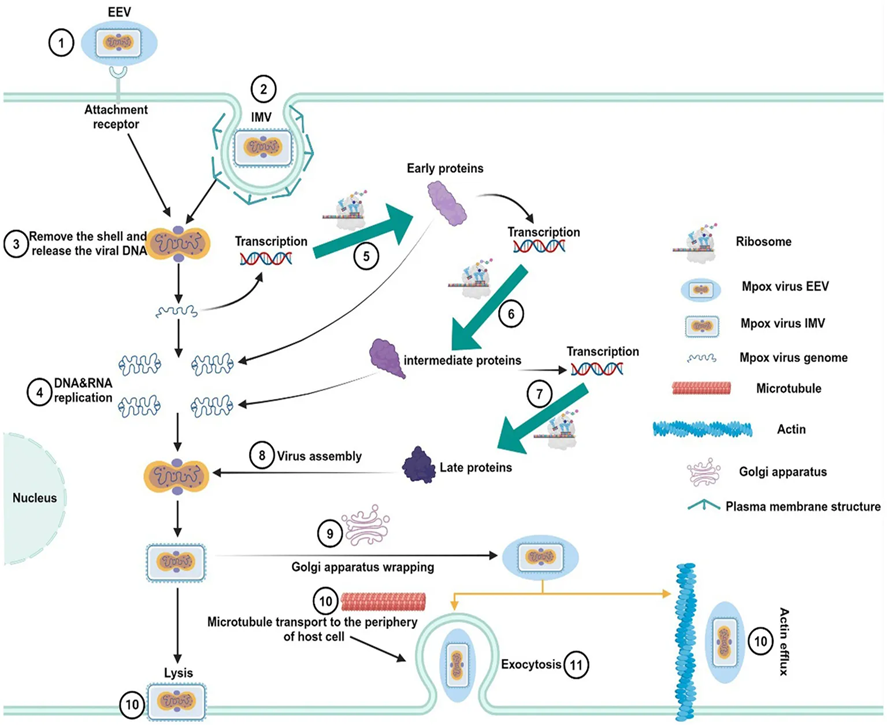
Fig 4: Replication life cycle of MPXV (Chen et al., 2025)
Entry, Uncoating, and Factory Formation
- Attachment and entry
- MV and EV forms attach via multiple viral proteins to ubiquitous cell surface molecules (e.g., glycosaminoglycans).
- Fusion at the plasma membrane or within endosomes delivers the viral core into the cytoplasm.
- Early transcription and partial uncoating
- Core carries RNA polymerase, transcription factors, and capping enzymes; early genes are transcribed immediately upon entry.
- Early products:
- Proteins that drive further uncoating.
- DNA replication machinery (E9, D4/E4, D5, A20/A22, I3 ssDNA-binding protein, B1 kinase).
- Innate immune antagonists.
- Viral factory (virosome) formation
- Endoplasmic reticulum (ER) membranes are reorganized around DNA to create a protected replication compartment.
- Viral factories concentrate replication and assembly proteins and shield DNA from cytoplasmic DNA sensors.
Initiation of DNA Replication
- Origin and hairpin structure
- Replication likely initiates via a nick near one telomeric hairpin; strand displacement from hairpin resembles the parvovirus-like model.
- Hairpin mismatches and loops are critical for proper genome maturation and packaging; altering the mismatch number impairs virion assembly without blocking DNA synthesis.
- Core replication machinery
- E9: high-fidelity B-family DNA polymerase.
- A22–A20 (MPXV F8-A22-E4 complex): processivity cofactor; in MPXV, A22-E4 undergoes state-dependent conformational changes affecting proofreading.
- D5: primase–helicase; unwinds DNA and synthesizes RNA primers.
- I3: ssDNA-binding protein stabilizing single-stranded regions.
- D4/E4: uracil-DNA glycosylase coupling base excision repair with replication.
- B1: Ser/Thr kinase regulating replication and repair steps.
- Mechanism (updated structural insight)
- Cryo-EM of MPXV polymerase (F8–A22–E4) shows:
- Polymerase transitions between replication and editing (proofreading) states, comparable to cellular B-family polymerases.
- F8 catalytic subunit carries polymerase and 3′–5′ exonuclease domains for mismatch removal.
- A22–E4 complex modulates DNA positioning; conformational shifts fine-tune switching to exonuclease site.
- E4 (D4 ortholog) not only supports replication but also functions as uracil-DNA glycosylase, initiating BER and directly linking lesion repair to ongoing replication.
- Cryo-EM of MPXV polymerase (F8–A22–E4) shows:
Elongation and Concatemer Formation
- Strand displacement synthesis
- Once initiated, replication produces long concatemers: alternating head-to-head and tail-to-tail repeats of the genome (LL and RR junctions).
- Newly synthesized DNA in factories is predominantly concatemeric, not unit length.
- Homologous recombination and repair
- Replication is tightly coupled to a high-frequency homologous recombination system.
- D4/E4, D5, A20/A22, I3, B1, and a Holliday junction endonuclease collectively support repair of stalled forks, resolution of concatemers, and generation of terminal ITRs.
- This replication-linked recombination also drives gene duplications, deletions, and mosaic genomes, as seen in recombinant cowpox CPXV-No-H2 and other orthopoxviruses.
Resolution to Unit Genomes and Telomere Processing
- Concatemer resolution
- Virus-encoded Holliday junction resolvase cleaves concatemers to yield unit-length genomes bearing correct hairpin termini.
- Internal deletions within concatemers before resolution can create expanded ITRs and duplicated terminal genes.
- Functional role of hairpin mismatches
- Experimental alteration of vaccinia telomeric mismatches shows:
- Viruses with heavily reduced mismatches exhibit impaired maturation, accumulating at the immature virion stage.
- DNA synthesis and telomere resolution remain intact; the main defect lies in genome packaging and core assembly.
- Indicates that hairpin secondary structure and mismatches are recognized by DNA-binding proteins (e.g., I1) during assembly.
- Experimental alteration of vaccinia telomeric mismatches shows:
Late Gene Expression and Virion Assembly (Replication–Assembly Interface)
- Transition from replication to late expression
- When sufficient DNA is synthesized, the transcription program shifts to late genes, encoding:
- Major core proteins (e.g., A3, A10, L4).
- Scaffold protein D13, membrane proteins, proteases.
- Enzymes to be packaged into virions (RNA polymerase, transcription factors).
- When sufficient DNA is synthesized, the transcription program shifts to late genes, encoding:
- Immature virion (IV) formation
- Crescent membranes form within factories, stabilized externally by D13 trimers.
- Membrane source is remodeled ER, dependent on L2 and other assembly factors.
- Crescents close to spherical IV, which accumulate core proteins and unit-length genomes.
- Maturation to IMV and wrapped forms
- Proteolytic processing of core proteins and structural reorganization convert IV → mature IMV.
- Some IMV acquire additional membranes from trans-Golgi or endosomes to form IEV → CEV → EEV, underpinning cell-to-cell spread and systemic dissemination.
Fidelity, Mutation, and Genome Plasticity
- High intrinsic fidelity with targeted flexibility
- Orthopox DNA polymerase (E9/F8) exhibits high fidelity with proofreading exonuclease activity, explaining low point mutation rates compared to RNA viruses.
- Structural data show a well-defined editing pathway for mismatched nucleotides, maintaining genome integrity.
- Recombination and “genomic accordion.”
- Despite low substitution rates, recombination, gene gain/loss, and terminal expansions/contractions generate significant genomic diversity.
- LCRs, tandem repeats, and terminal gene duplications can expand or contract rapidly under selection (e.g., host jumps, immune pressure).
- This “accordion” mechanism allows quick modulation of host-interaction genes (e.g., OPG153, OPG204, OPG208, OPG027, D1L) while preserving core replication machinery
Pathogenesis and Host Immune Response
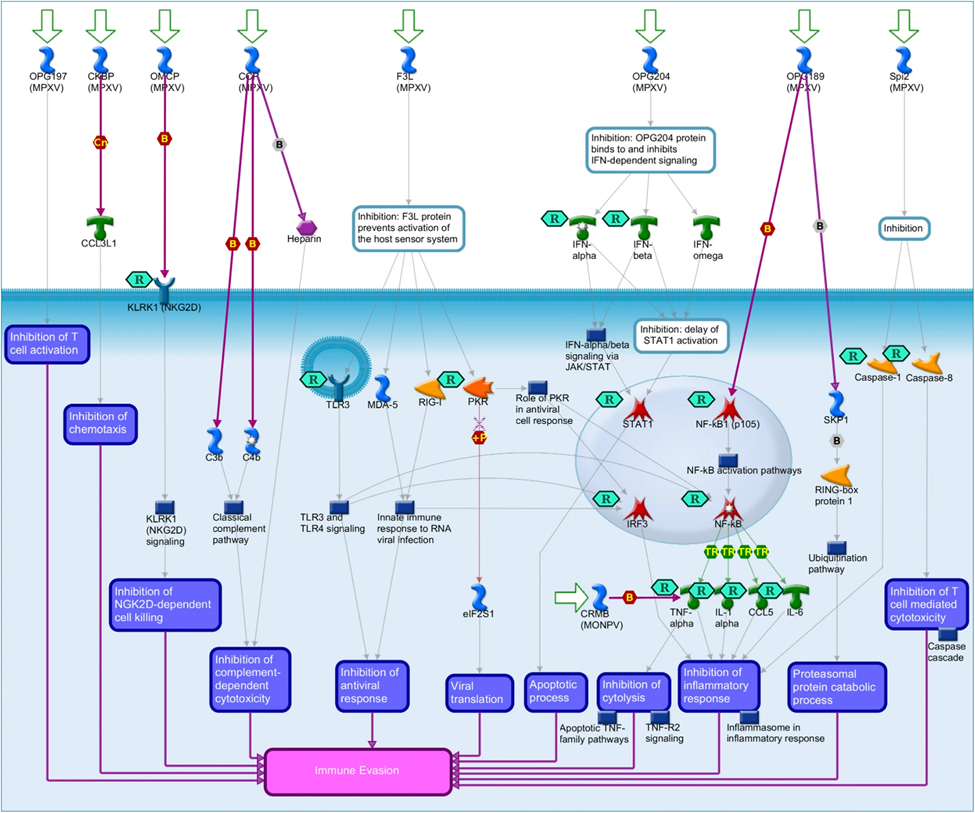
Fig 5: Mpox immune evasion pathway map (Hajjo et al., 2025)
- Entry and primary replication sites
- Inoculation via skin/mucosa, respiratory tract, or genital/anal mucosa (notably in mpox 2022–25).
- Initial replication in local epithelial cells/dendritic cells → spread to regional lymph nodes → viremia → systemic dissemination, especially to skin and mucosal epithelium.
- Tissue targets
- Skin, oropharyngeal and anogenital mucosa, respiratory epithelium; in severe mpox, lung, GI tract, and CNS can be involved.
- Immune response
- Robust innate response: type I IFN, NK cells, macrophages; heavily countered by viral IFN/TNF/chemokine-binding proteins.
- Strong humoral immunity: neutralizing antibodies to core and envelope proteins; basis of cross-protection between orthopoxviruses.
- Cell-mediated immunity (CD4+, CD8+ T cells) is critical for viral clearance and long-term protection; severe/prolonged disease in advanced HIV and other immunocompromised hosts.
- Immune evasion strategies
- Secreted cytokine/chemokine decoys.
- Inhibitors of PRR signaling (e.g., TLR, STING, NF-κB, IRF pathways).
- Blockade of apoptosis and antigen presentation.
- Neurological and systemic complications in mpox (recent evidence)
- Common: headache (~29%), myalgia (~30%), fatigue (~33%).
- Rare but severe: encephalitis, encephalomyelitis, transverse myelitis, Guillain–Barré syndrome, Parsonage–Turner syndrome, autonomic dysfunction.
- Recovery variable; some patients have persistent deficits up to ≥32 weeks.
Epidemiology and Transmission
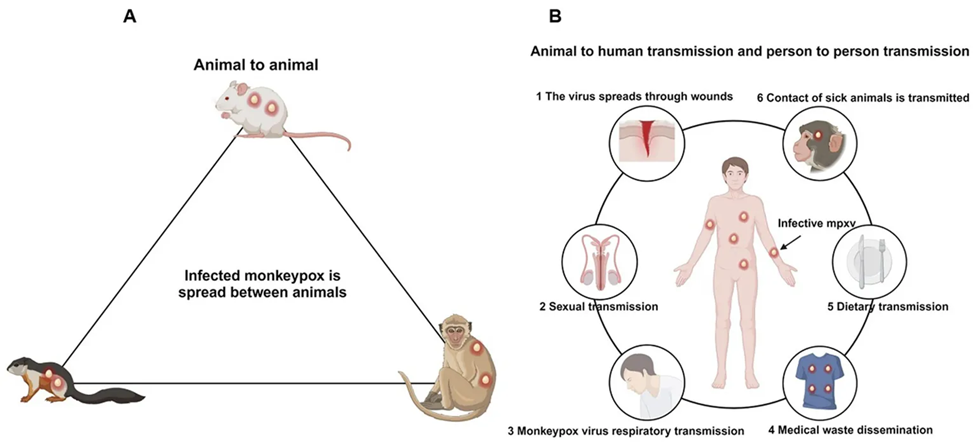
Fig 6: Transmission pathway of Mpox (Chen et al., 2025)
- Reservoirs and hosts
- Natural reservoirs: small mammals (especially rodents) for many orthopoxviruses; humans are the only reservoir for VARV.
- Classical transmission routes
- Zoonotic: contact with infected animals or animal products.
- Human-to-human:
- Close skin-to-skin contact with lesions or body fluids.
- Large respiratory droplets with prolonged face-to-face contact.
- Fomites (bedding, clothing).
- Mpox 2022–25 global outbreaks
- Shift from sporadic zoonotic cases in Central/West Africa to sustained human-to-human transmission in non-endemic countries.
- Dominant clade IIb B.1 lineage with enhanced human transmissibility.
- High proportion of cases among men who have sex with men (MSM), with transmission strongly associated with sexual networks and anal-genital contact.
- WHO designation as a Public Health Emergency of International Concern (PHEIC) in 2022.
- Drivers of re-emergence
- Waning population-level immunity after cessation of smallpox vaccination.
- Urbanization and increased human–animal interface in endemic regions.
- Viral evolution with gene loss and point mutations facilitating human-to-human spread.
- Immunocompromised populations (HIV, advanced AIDS) are experiencing more severe, prolonged disease.
Clinical Manifestations
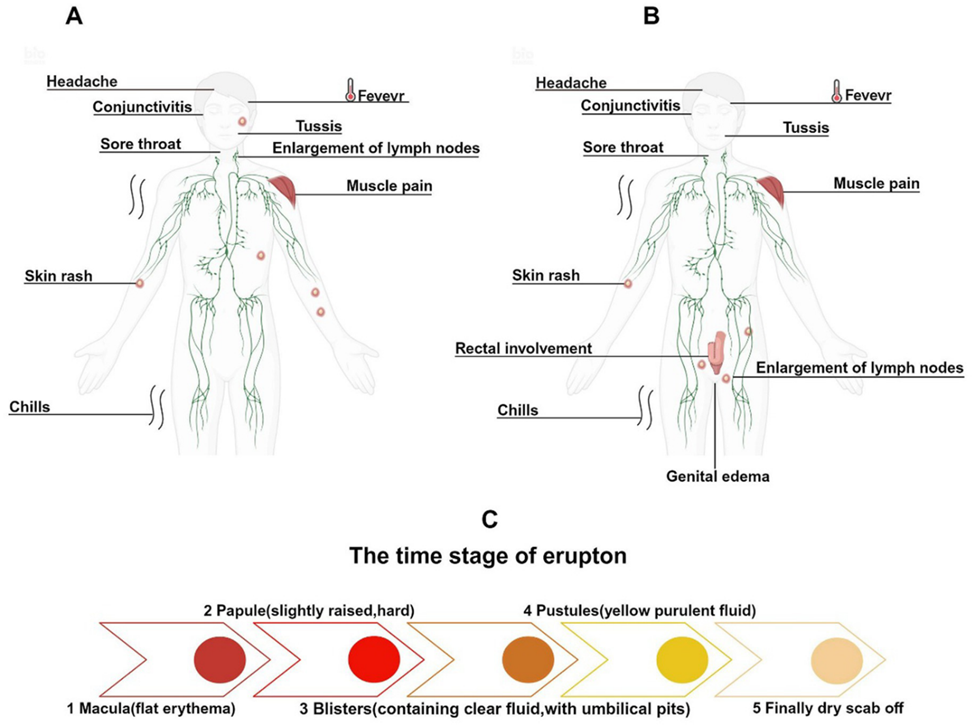
Fig 7: Clinical symptoms and rash evolution of Mpox infections across different historical periods (Chen et al., 2025)
Orthopoxvirus infections share a prototypical systemic illness with rash, but with virus- and host-specific nuances.
- General mpox/orthopox clinical pattern:
- Incubation: ~5–21 days.
- Prodrome:
- Fever, chills, malaise.
- Marked lymphadenopathy (distinguishes mpox from smallpox).
- Myalgia, headache, back pain.
- Rash:
- Progression from macules → papules → vesicles → pustules → crusts.
- Historically centrifugal distribution (face, extremities, including palms/soles).
- Duration: illness typically 2–4 weeks.
- Mpox in 2022+ outbreaks (clade IIb)
- Lesions are often localized to the genital, perianal, and oral regions; fewer, asynchronous lesions compared with classical descriptions.
- Prominent anogenital pain, proctitis, rectal bleeding, dyschezia.
- Gastrointestinal (GI) manifestations
- Proctitis, rectal pain, tenesmus, rectal bleeding.
- Nausea, vomiting, diarrhea, abdominal pain.
- Sexual transmission via receptive anal intercourse is associated with proctitis.
- Neurological manifestations
- Common: headache, myalgia, fatigue.
- Uncommon but serious: encephalitis, encephalomyelitis, transverse myelitis, Guillain–Barré syndrome; autonomic dysfunction (bowel/bladder incontinence).
- Complications
- Secondary bacterial skin infections, sepsis.
- Keratitis and vision loss.
- Pneumonia.
- In immunocompromised individuals: extensive necrotic lesions, prolonged viremia, multi-organ involvement, and higher mortality.
Laboratory Diagnosis
- Preferred specimens
- Skin lesion material: swabs of vesicular/pustular fluid or crusts.
- Oropharyngeal and anogenital swabs for mucosal disease.
- Blood for severe or disseminated disease.
- Primary diagnostic methods
- Real-time PCR (qPCR) targeting conserved orthopox and MPXV-specific genes → gold standard.
- High sensitivity; can be multiplexed with other rash-causing viruses.
- Supplementary techniques
- Conventional PCR and sequencing for clade/lineage assignment and genomic surveillance.
- Virus isolation in BSL-3 facilities for research; not routine.
- Electron microscopy: brick-shaped orthopox virions, but not species-specific.
- Serology (ELISA, neutralization): useful for exposure and immunity studies; limited for acute diagnosis due to cross-reactivity between orthopoxviruses and prior smallpox vaccination.
- Emerging tools
- Whole-genome sequencing for real-time molecular epidemiology and detection of adaptive mutations (e.g., OPG027, D1L changes).
- AI-assisted genomic surveillance pipelines to track variants and predict virulence.
Treatment and Antiviral Therapy
There is no orthopoxvirus-specific small-molecule fully validated in randomized mpox trials, but several repurposed antivirals and supportive strategies are used.
- First-line antiviral: Tecovirimat (TPOXX; ST-246)
- Targets VP37 (envelope protein) and blocks formation of EEV, limiting dissemination.
- Favored for moderate–severe disease, high-risk patients (e.g., advanced HIV, pregnant people, extensive skin disease).
- Other antivirals
- Cidofovir: nucleotide analogue inhibiting viral DNA polymerase; nephrotoxic, IV administration.
- Brincidofovir (Tembexa): oral lipid conjugate of cidofovir with improved safety profile.
- Ribavirin and other candidates with in vitro or animal efficacy, but limited clinical data.
- Newer candidates targeting thymidine kinase, VP39, and other essential enzymes under investigation via in silico and in vitro screens.
- Immunomodulatory/supportive therapy
- Vaccinia immune globulin intravenous (VIG-IV) for severe or vaccine-related orthopox complications, extrapolated to mpox in some settings.
- Analgesia, management of secondary bacterial infections, hydration, and nutritional support.
- In severe neurological mpox, corticosteroids and IVIG were used in case series.
Prevention and Control
- Vaccination
- Smallpox vaccines provide cross-protection against mpox and other orthopoxviruses due to shared antigens.
- Current vaccines used for mpox prevention:
- JYNNEOS/Imvamune/Imvanex (MVA-BN; non-replicating modified vaccinia Ankara).
- ACAM2000 (replicating vaccinia-based vaccine; greater adverse-event profile).
- Strategies: pre-exposure prophylaxis for high-risk groups (e.g., MSM with multiple partners, laboratory personnel), and post-exposure prophylaxis for close contacts, ideally within 4 days of exposure.
- Non-pharmaceutical interventions
- Isolation of symptomatic cases until lesions crust and fall off.
- Contact tracing and ring vaccination where vaccines are available.
- Use of personal protective equipment (PPE) in healthcare and household settings.
- Environmental decontamination of fomites (bedding, clothing, surfaces).
- Public-health challenges and priorities
- Global inequities in vaccine and antiviral access, particularly in African endemic regions.
- Post-smallpox-eradication immunity gap; large unvaccinated younger cohorts.
- Need for strengthened genomic surveillance to detect emergent clades/lineages with altered virulence or transmission.
- Stigma in affected sexual networks is hindering case finding and adherence to isolation.
Conclusion
Orthopoxviruses are a paradigmatic group of large, cytoplasmic DNA viruses with complex genomes that encode self-contained replication and immune-evasion machinery. The resurgence of mpox since 2022 has underscored the continued relevance of orthopoxvirus biology, particularly in genomic evolution, immune modulation, and human-to-human transmission. Advances in molecular virology, sequencing-based surveillance, and cross-protective vaccination provide powerful tools to contain current and future outbreaks, but gaps persist in equitable access, targeted antivirals, and understanding of long-term complications. Sustained research and integrated public-health strategies are essential to manage the ongoing and potential future burden of orthopoxvirus infections.
References
- Karagoz, A., Tombuloglu, H., Alsaeed, M., Tombuloglu, G., AlRubaish, A. A., Mahmoud, A., … Alsuhaimi, E. (2023). Monkeypox (mpox) virus: Classification, origin, transmission, genome organization, antiviral drugs, and molecular diagnosis. Journal of Infection and Public Health, 16(2), 265–279. https://doi.org/10.1016/j.jiph.2022.12.010
- Bhandari, G., Acharya, A., Chettri, A. K., & Sharma, S. (2025). Neurological manifestations of Mpox virus during the recent global outbreak: A systematic review. BMC Infectious Diseases, 25, Article 812. https://doi.org/10.1186/s12879-025-10361-0
- Obi, C. L., Mkolo, N., Mugivhisa, L. L., Ogunrombi, M. O., Mphephu, M. M., & Naidoo, C. (2025). Mpox: An emerging or re-emerging infection with a potential colossal burden on healthcare globally. African Journal of Laboratory Medicine, 14(1), a2332. https://doi.org/10.4102/ajlm.v14i1.2332
- Paulino-Moreno, Z., & Gutiérrez-Pérez, B. (2025). Emerging technologies in genomic surveillance and control of the Mpox virus: Artificial intelligence and sequencing for epidemiological control—A systematic review. International Journal of Educational Practices and Engineering, 5(1), 1–20. (check article for final DOI)
- Cambaza, E. M. (2025). A review of the molecular understanding of the Mpox virus (MPXV): Genomics, immune evasion, and therapeutic targets. Zoonotic Diseases, 5(1), 3. https://doi.org/10.3390/zoonoticdis5010003
- Ramakrishnan, R., Shenoy, A., Madhavan, R., & Meyer, D. (2024). Mpox gastrointestinal manifestations: A systematic review. BMJ Open Gastroenterology, 11(1), e001355. https://doi.org/10.1136/bmjgast-2023-001355
- Ru, E., & Ogbodo, E. C. (2025). Monkeypox; pathogenesis, risk factors, complications, laboratory diagnosis, prevention and management: A concise review. Saudi Journal of Medicine, 10(7), 312–324. https://doi.org/10.36348/sjm.2025.v10i07.004
- Babkin, I. V., Babkina, I. N., & Tikunova, N. V. (2022). An update of Orthopoxvirus molecular evolution. Viruses, 14(2), 388. https://doi.org/10.3390/v14020388
- Krumova, S., Ivanov, D., & Christova, I. (2025). A review of genome organization, evolution, transmission, circulation, and clinical manifestation of monkeypox virus. Problems of Infectious and Parasitic Diseases, 53(1), 5–20. (check article for final DOI)
- Rohaim, M. A., Naggar, R. F. E., Atasoy, M. O., & Munir, M. (2024). Molecular virology of orthopoxviruses with special reference to monkeypox virus. In Advances in Experimental Medicine and Biology (Vol. 1451, pp. 111–124). Springer. https://doi.org/10.1007/978-3-031-39651-2_6
- Hajjo, R., Abusara, O. H., Sabbah, D., & Bardaweel, S. (2025). Advancing the understanding and management of Mpox: Insights into epidemiology, disease pathways, prevention, and therapeutic strategies. BMC Infectious Diseases, 25, Article 312. https://doi.org/10.1186/s12879-025-09584-x
- Chen, S., Huang, J., Chen, J., Liu, F., Wang, S., Wang, N., … Liu, Z. (2025). Mpox virus: Virology, molecular epidemiology, and global public health challenges. Frontiers in Microbiology, 16, 1507694. https://doi.org/10.3389/fmicb.2025.1507694
- Begum, J. P. S., Ngangom, L., Semwal, P., Painuli, S., & Sharma, R. (2023). Emergence of monkeypox: A worldwide public health crisis. Human Cell, 36(1), 1–18. https://doi.org/10.1007/s13577-022-00784-3
- Mitjà, O., Ogoina, D., Titanji, B. K., Galvan, C., Muyembe, J. J., & White, R. G., et al. (2023). Monkeypox. The Lancet, 401(10370), 60–74. https://doi.org/10.1016/S0140-6736(22)02075-X
- Sklenovská, N., & Van Ranst, M. (2018). Emergence of monkeypox as the most important orthopoxvirus infection in humans. Frontiers in Public Health, 6, 241. https://doi.org/10.3389/fpubh.2018.00241
- Alakunle, E., Moens, U., Nchinda, G., & Okeke, M. I. (2020). Monkeypox virus in Nigeria: Infection biology, epidemiology, and evolution. Viruses, 12(11), 1257. https://doi.org/10.3390/v12111257
- Farias, L. A. B. G., et al. (2023). Severe mpox in patients with advanced AIDS: Long-term disease and fatal outcome. Revista da Sociedade Brasileira de Medicina Tropical, 56, e0228-2023. https://doi.org/10.1590/0037-8682-0228-2023
- Siegrist, C. M., Kinahan, S. M., Settecerri, T., Greene, A. C., & Santarpia, J. L. (2020). CRISPR/Cas9 as an antiviral against Orthopoxviruses using an AAV vector. Scientific Reports, 10(1), 19499. https://doi.org/10.1038/s41598-020-76449-9
- Lansiaux, E., Jain, N., Laivacuma, S., & Reinis, A. (2022). The virology of human monkeypox virus (hMPXV): A brief overview. Virus Research, 322, 198932. https://doi.org/10.1016/j.virusres.2022.198932
