In recombinant DNA technology (RDT), the terminology vector refers to a carrier that allows the transfer of genetic material from one cell to another or from one organism to another. A vector or carrier is required to safely transport the foreign genetic material, deliver it, and make it functional in the host cell.

A plasmid is an extrachromosomal, circular, and self-replicating ds-DNA molecule predominantly present in bacteria. In nature, plasmids carry accessory genetic elements that confer advantages under specific conditions, such as antibiotic resistance, pathogenesis, and metabolism. Naturally occurring plasmids are changed or altered (size reduction or addition of desired sequences) in the laboratory so that they can deliver, multiply, and express foreign genetic material into the host cells. Plasmids’ size can range from 1kb to up to 500kb and can carry DNA inserts up to 10kb.
Essential Components of Plasmids
Most plasmid vectors contain the same functional elements. The basic components of a plasmid vector contain the Origin of Replication (ORI), selection or marker gene, multiple cloning site (MCS), and regulatory sequences (promoters/terminators). Additionally, plasmids also contain sites for gene inserts and primer binding (for initiation and termination during PCR).
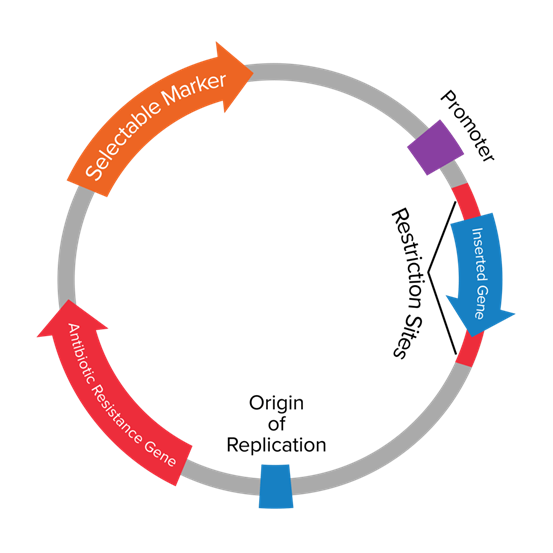
The Origin of Replication (ORI) of Plasmids
The Origin of Replication (ori) is a DNA sequence on a plasmid where DNA replication initiates, allowing the plasmid to replicate within the host cell. This site provides assembly sites for different components of host replication machinery, as the plasmid replication depends upon the host machinery and host enzymes. The ORI site is rich in A-T pairs, as these sites can be separated more easily at a lower temperature than G-C pairs.
Host Range: Sometimes, two or three ORI sites could also be present to be exclusively recognized by broad hosts, thus enabling replication across diverse bacteria. Similarly, a narrow host range maintains and allows plasmid replication in closely related species only.
Replication in plasmids: Plasmids can replicate autonomously without depending upon the host cell (however, they utilize the host cell components: polymerases, helicases, dNTPs) and follow the rolling circle mechanism.
Selectable Markers of Plasmids
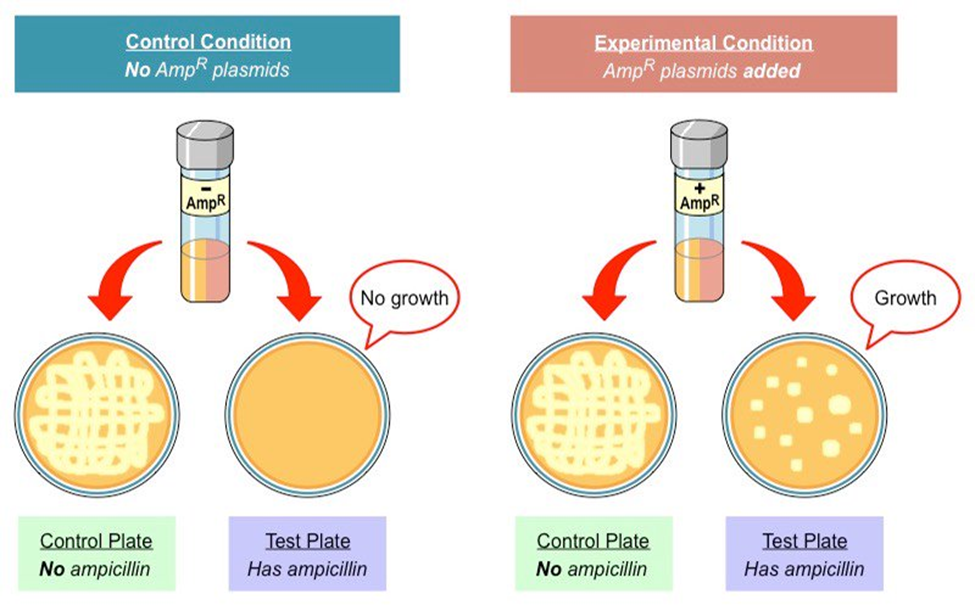
A selectable marker in plasmids is used to identify cells that successfully take up the plasmid. To distinguish transformants from non-transformants (the cells that did not receive the plasmid), one of the most preferred techniques is the use of antibiotic-resistant genes as selectable markers. For instance, when bacteria are grown in a medium containing an antibiotic, only the cells carrying the plasmid with the resistant genes can survive and grow; cells lacking the plasmid cannot grow and are hence eliminated by the antibiotic. Selective markers, therefore, make sure that plasmids are maintained during cellular replication and growth to obtain the desired trait.
Ampicillin (AmpR), kanamycin (KanR), and tetracycline resistance gene (TetR) are some of the antibiotic resistance genes used as markers in plasmids. Additionally, genes coding for fluorescent proteins, such as Green Fluorescent Protein (GFP), are also used as markers to visually identify transformants based on fluorescence.
The Multiple Cloning Site (MCS) of Plasmids
A multiple cloning site (MCS) or a polylinker is a short region of DNA in a plasmid that contains multiple restriction sites (sequence recognition sites for various restriction enzymes). As the restriction enzymes cleave ds-DNA at specific nucleotide sequences, the presence of multiple restriction sites within the MCS provides greater flexibility for foreign DNA insertion into the plasmid. The MCS may contain up to 20 restriction sites in an engineered plasmid.
During recombination, the plasmid DNA is cleaved at a selected restriction site within the MCS. This creates an opening for a gene of interest (GOI) to be inserted by DNA ligase-mediated ligation. Here, both the vector DNA and the GOI are cut with the same restriction enzyme to make their ends compatible with each other. MCSs are designed in such a way that their sequences do not appear in other vector elements, or GOI, to avoid undesired cleavage. After ligation, a recombinant plasmid vector containing the target gene is formed.
Promoter and Terminator Sequences of Plasmids
Promoters are typically the binding site of transcription factors, such as enzymes (RNA polymerases) and regulatory proteins, to control transcription (gene expression). Promoters can be as short as a few nucleotides to hundreds of nucleotides long. They are non-coding sequences located upstream (5’ region) of the target DNA. The binding of the host RNA polymerase to a promoter region in the plasmid initiates transcription of mRNA, therefore regulating the gene expression of the recombinant gene. Common promoters used in plasmid vectors include the lac promoter (lactose or IPTG inducible).
Terminators are also the non-coding region of a gene located downstream (3’ region) of a target gene. In plasmid vectors, termination occurs in a rho-independent manner. The terminator region of the plasmid contains GC-rich inverted repeats, which cause hairpin loops to form in the subsequently formed mRNA. This causes RNA polymerase to stop transcribing, thereby releasing the formed mRNA.
Cloning vs. Expression Vectors of Plasmids
Based on practical implications, either cloning or expression vector is used in RDT. Both cloning and expression vectors function as carriers that introduce target DNA into a host cell. Generally, these vectors share core structural elements, such as ORI sites, restriction sites, and selectable markers, but differ in function.
Cloning vectors are designed to transport, maintain, and amplify target DNA into a host. These vectors prioritize amplification of target DNA rather than its expression. Cloning vectors are optimized for stability and easy manipulation, which makes the insertion of a genetic element efficient and straightforward. If the objective is to amplify a target gene, sequence a gene fragment, or build a DNA library, a cloning vector is appropriate. Plasmids, bacteriophages, or Cosmids are used as cloning vectors. The choice of a cloning vector depends mainly upon the size of the DNA insert.
Expression vectors, mostly plasmids, are specialized types of cloning vectors. While cloning vectors mainly function to amplify target DNA sequences into the host, expression vectors are designed to specifically produce RNA and proteins from the target DNA. An expression vector, in this case, utilizes the host’s transcriptional and translational mechanisms to synthesize the target mRNA and proteins, respectively. Additional elements, such as promoters, terminators, ribosome binding sites, transcription and translation initiation sites, etc., are present. Expression vectors also contain fusion tags, such as His-tags, for efficient detection of the recombinant protein.
Copy Number Control of Plasmids
Plasmid copy number (PCN) refers to the number of plasmids present in a single cell. PCN varies broadly, ranging from a single plasmid to more than 1000 in a cell. An appropriate PCN is essential in a cell to maintain plasmid stability and long-term survival. Too high PCN can cause a significant metabolic burden to the host cell by consuming its resources, whereas too low a copy number can cause a risk of lacking inheritance during cell division.
Plasmid copy number (PCN) is controlled by many regulatory factors present in the plasmid as well as limitations set by the host cell. Some of the factors responsible are:
Replication control: TheORI site and its initiator proteins regulate how plasmid replication initiates, and small changes in these components directly increase or decrease PCN.
Antisense RNA: Small RNAs, such as RNA-I, complementary to essential RNAs, inhibit excessive replication.
Repressor proteins: Proteins such as Rop (repressor of primer) can enhance the binding of antisense RNA to target DNA, further suppressing replication.
Partitioning systems: In low-copy-number plasmids, proteins such as ParA and ParB help distribute plasmid copies evenly to daughter cells during cell division, such that each daughter cell receives at least one copy.
Plasmid size: Generally, smaller plasmids have high copy numbers, whereas larger plasmids have lower PCN due to increased metabolic demands in the host.
Toxin-antitoxin systems: After host cell replication, when the daughter cell loses the plasmid, the unstable antitoxin degrades, allowing the toxin to function and kill the cell. This ensures that only plasmid-bearing cells survive.
Common Plasmid Examples (pBR322, pUC19)
pBR322
pBR322 plasmid, derived from pBR313 and developed from E. coli, was constructed by Francisco Bolivar and team in the late 1970s at the University of California. It is one of the most widely used vectors in molecular biology. The vector is named after its creators: Francisco Bolivar Zapata and Raymond L. Rodriguez; p represents the plasmid, and 322 represents the clone number of the vector.
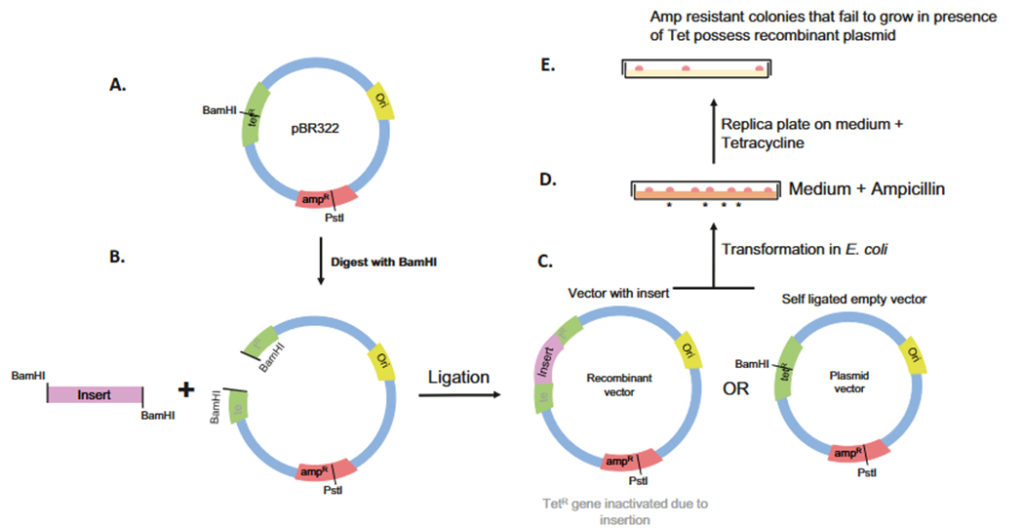
pBR322 is a 4362-bp ds-DNA plasmid cloning vector, which contains an ORI site, carries ampicillin (AmpR) and tetracycline resistance (TetR) genes as markers, and multiple restriction sites for restriction enzymes: PstI, EcoRI, HindIII, BamHI, and SalI. The ORI site is derived from ColE1, which ensures 15-20 PCN.
The selection method starts with the insertion of the target DNA at the BamHI restriction site within the TetR gene. The TetR gene then becomes inactivated due to the insertion of the target DNA. Here, the selection markers work in two steps: positive and negative selection. When transformed bacterial cells are first grown in ampicillin-containing media, all of the cells containing plasmid can grow due to the presence of the AmpR gene (positive selection). However, when these viable colonies are cultured on tetracycline-containing media, only plasmids with the TetR gene grow, eliminating recombinant colonies (negative selection). Therefore, the desired colonies that show ampicillin resistance, but tetracycline sensitivity show that they carry the recombinant plasmid.
pUC19
The pUC vector is a better version of the pBR322 plasmid. It was developed using the components from pBR322, also at the University of California. The positive-negative screening method in pBR322 was relatively time-consuming and often led to errors during selection. pUC vectors were then introduced with a single-step screening process called the blue-white screening to overcome the limitations of pBR322. The most common types among the pUC series are pUC18 and pUC19.
The 2686-bp-long pUC19 vector contains the AmpR and lacZα gene that codes for the alpha peptide of the β-galactosidase enzyme, for the selection of transformants. The selection through AmpR occurs in the same fashion as the positive selection mechanism in the pBR322 plasmid.
The principle in selection during transformation of pUC19 vectors by LacZα-basedis mainly dependent upon the complementation of alpha peptide and omega peptide to produce a functional β-galactosidase enzyme. Selection is initiated after the insertion of the target DNA into a restriction site within the MCS. As the MCS is within the coding region of the lacZα gene, this insertion disrupts the lacZα gene. This process is called insertional inactivation. Then, the transformed bacteria are cultured on a chromogenic medium containing substrate for β-galactosidase. Recombinants (containing target DNA that disrupts the lacZα gene) do not produce the enzyme and appear white, whereas plasmids without the target DNA disrupt the coding region of the lacZα gene, fail to produce the enzyme, and appear white. Therefore, blue colonies represent non-recombinant plasmids, whereas white colonies represent recombinant ones. This overall process is referred to as blue-white screening.
Applications of Plasmids in Genetic Engineering
- Construction of rDNA: In gene cloning, specific target DNA is inserted into plasmids to generate recombinant DNA (rDNA), which allows for obtaining large quantities of target DNA for downstream analyses, such as genetic analysis, sequencing, or gene manipulation.
- Recombinant protein production: Plasmid-based expression vectors are commonly used to produce recombinant proteins, such as vaccines, in host cells, allowing their expanded use in research, medicine, and biotechnology.
- Metabolic engineering: Due to their rapid metabolism and industrial relevance, plasmid vectors are highly applied to introduce targeted genetic changes, such as insertions and deletions, to optimize metabolic flux and product yield in microorganisms, especially bacteria.
- Plant genetic engineering: Engineered plasmid vectors are used to deliver gene editing constructs into plant genomes for crop improvement by generating tolerance against various biotic and abiotic factors.
Conclusion
In conclusion, plasmid vectors, due to their simple structure and ability to replicate independently in the host, are essential in rDNA technology and genetic engineering. Plasmids can be tailored to different applications, extending to microbial, plant, and animal systems, hence making them a versatile tool in biotechnology research. Current advancements in molecular biology techniques further enhance plasmids and plasmid-based technology to develop effective solutions.
References
- Patil R., Sivaram A., Patil N. In: A complete guide to gene cloning: from basic to advanced. Patil N., Sivaram A., editors. Springer; Cham, Switzerland: 2022. Gene isolation methods: beginner’s guide
- https://blog.addgene.org/plasmids-101-what-is-a-plasmid
- Lloyd, G. S., & Thomas, C. M. (2023). Microbial Primer: The logic of bacterial plasmids. Microbiology (Reading, England), 169(7), 001336. https://doi.org/10.1099/mic.0.001336
- https://blog.addgene.org/plasmid-101-origin-of-replication
- Douglas F. Browning, Rita E. Godfrey, Kirsty L. Richards, Colin Robinson, Stephen J.W. Busby; Exploitation of the Escherichia coli lac operon promoter for controlled recombinant protein production. Biochem Soc Trans 30 April 2019; 47 (2): 755–763. doi: https://doi.org/10.1042/BST20190059
- https://blog.addgene.org/plasmids-101-terminators-and-polya-signals
- Furuya, N., & Komano, T. (2000). Initiation and termination of DNA transfer during conjugation of IncI1 plasmid R64: roles of two sets of inverted repeat sequences within oriT in termination of R64 transfer. Journal of bacteriology, 182(11), 3191–3196. https://doi.org/10.1128/JB.182.11.3191-3196.2000
- https://pediaa.com/difference-between-cloning-vector-and-expression-vector/
- Basanta Kumara Behera, Chapter 3 – Biological systems for therapeutic protein production, In Developments in Applied Microbiology and Biotechnology, Conceptual Development of Industrial Biotechnology for Commercial Production of Vaccines and Biopharmaceuticals, Academic Press, 2023, Pages 65-99, ISBN 9780443187483, https://doi.org/10.1016/B978-0-443-18748-3.00004-4.
- https://en.wikipedia.org/wiki/Plasmid_copy_number
- Ramiro-Martínez, P., de Quinto, I., Lanza, V.F. et al. Universal rules govern plasmid copy number. Nat Commun 16, 6022 (2025). https://doi.org/10.1038/s41467-025-61202-5
- del Solar, G., & Espinosa, M. (2000). Plasmid copy number control: an ever-growing story. Molecular microbiology, 37(3), 492–500. https://doi.org/10.1046/j.1365-2958.2000.02005.x
- Bolivar, F., Rodriguez, R. L., Greene, P. J., Betlach, M. C., Heyneker, H. L., Boyer, H. W., Crosa, J. H., & Falkow, S. (1977). Construction and characterization of new cloning vehicles. II. A multipurpose cloning system. Gene, 2(2), 95–113.
- Leonard G. Davis, Mark D. Dibner, James F. Battey, SECTION 4-1 – pBR322, Editor(s): Leonard G. Davis, Mark D. Dibner, James F. Battey, Basic Methods in Molecular Biology, Elsevier, 1986, Pages 24-25, ISBN 9780444010827, https://doi.org/10.1016/B978-0-444-01082-7.50010-2.
- RNA-Guided Recombinase-Cas9 Fusion Targets Genomic DNA Deletion and Integration. Standage-Beier K, Brookhouser N, Balachandran P, Zhang Q, Brafman DA, Wang X. CRISPR J. 2019 Aug;2:209-222. doi: 10.1089/crispr.2019.0013. PubMed 31436506
- Sokra, I., Meta, H., Lika, R., Socheata, C., & Somaly, S. (2026). Plasmid vectors for gene editing and genetic engineering: Design principles, efficiency, and applications. Journal of Agriculture and Technology, 2(1), 254-264.
